-
DNA是大多数生物的遗传信息载体,是保证物种延续性的基础,微小的变化都可能对细胞产生致命的影响[1]。在生命进化的过程中,DNA一直受到外界环境和内部因素的影响。细胞中dNTPs和rNTPs浓度改变、DNA聚合酶保真性变化和外界环境改变都可能导致高突变率和非完全同源DNA序列之间的重组能力增加,进而影响基因组的稳定性[2]。DNA错配修复(mismatch repair,MMR)系统可以纠正复制和重组过程中发生的DNA碱基错配,确保染色体复制的精确性和维持基因组的稳定性[3]。
MutL/MutS是原核生物最重要的MMR系统。DNA错配修复蛋白MutL羧基端含MLH1和PMS2二聚体结构域,其中PMS2具有内切酶活性;MutL氨基端具有ATP酶和DNA结合活性[4-6]。当MMR复合体中的另一个关键蛋白MutS识别并结合到发生碱基错配位点时,其构象发生变化,招募MutL形成修复复合体,对错配序列进行修复[7-8]。
在大肠杆菌中,MutL发挥内切酶作用,在错配位置产生切口[9],阻断DNA链继续合成。MutL的DNA结合位点突变会使MutL无法结合在发生错误的DNA位点,MMR系统不能发挥正常功能,活性下降,错配序列无法修复,显著提高细菌的突变率[10-12]。在铜绿假单胞菌中,MutL也具有DNA切割活性,使发生错配的DNA双链断裂,然后通过非同源端连接进行DNA修复。非同源端连接将提高染色体缺失的可能性,染色体的缺失导致铜绿假单胞菌产生棕色突变体,这种颜色的突变体可以防止噬菌体捕食[13]。在人和多数模式生物中,MutL的功能已有广泛研究[14-16],但其是否与病原菌的致病性相关还未见报道。
白叶枯病是影响水稻生产最严重的细菌性病害,由水稻黄单胞菌白叶枯致病变种(Xanthomonas oryzae pv. oryzae, Xoo)侵染所致[17]。Xoo致病力受很多因素调控,调控途径也不相同,不同途径间还有相互影响,是外界环境和内在因素共同作用的结果[18]。目前的研究表明,基础代谢、外源信号感应与传递、分泌系统及其分泌蛋白是调控Xoo致病力的关键因素[19-20]。Xoo侵染水稻时,水稻会产生防卫反应,产生大量有毒的物质如活性氧等毒害病原菌[18],导致其突变增加,生存受到挑战[19]。病原菌要成功侵染必须要有高效的应对措施。DNA损伤修复是保证病原菌应对外界环境不利影响的重要措施之一[9]。因此,MMR应该在此过程中起到非常重要的作用,但相关研究还未见报道。本实验研究MutL是否参与Xoo DNA错误修复以及是否调控病原菌的致病力,旨在为白叶枯病的防治提供一定的理论基础。
-
PXO99A 菌株和pK18mobSacB质粒为笔者所在的实验室保存,大肠杆菌菌株购自全式金生物技术有限公司。
-
PSA培养基:1%胰蛋白胨、1%蔗糖、0.1%谷氨酸钠;PGA培养基:1%胰蛋白胨、1%葡萄糖、0.1% 谷氨酸钠;PSSU培养基:1%胰蛋白胨、15%蔗糖、0.1%谷氨酸钠;LB培养基:1%胰蛋白胨、0. 5%酵母提取物、1% NaCl;M4M培养基:0.048% Na2HPO4、0.03% KH2PO4、0.05%柠檬酸钠、0.1% (NH4)2SO4、0.05%酶水解酪素、0.02% MgSO4·7H2O、0.05%葡萄糖。固体培养基另加1. 5%的琼脂。DNA凝胶回收及质粒提取试剂盒购自天根生化科技有限公司。高保真DNA聚合酶、限制性核酸内切酶及T4连接酶购自NEB公司。其他试剂购自广州化学试剂公司。
-
根据基因两端序列,运用软件Primer 5分析、设计引物(表1),并由生工生物工程(广州)分公司合成。
表 1 本实验引物
引物名 序列(5′-3′) 用途 产物大小/bp PX0410ddFF GTCGAAGCTTGCCAATGTGTTGCTCGACC mutL deletion 540 PX0410ddFR TAGCGGATCCTGACGGATCGCCATCAGCGC PX0410ddRF TAGCGGATCCGGCTCGGTTTACGCTGAGCG 572 PX0410ddRR CGTGGAATTCATGCTTGACTCGCAGCGCAT PX0410hmF CAGCAAGCTTGATGGCGATCCGTCAGTTGC ΔmutL complementation 1 918 PX0410hmR GAATTCAAGCGTAGTCTGGGACGTCGTATG
GGTAACGTCCCCTGAGAAACCAAC -
PCR 扩增mutL上下游各约500 bp序列,分别用HindIII/BamHI和BamHI/EcoRI酶切后一起连接到经HindIII/EcoRI酶切的自杀性载体pK18mobSacB上,热激法转化大肠杆菌,获得pK18-mutL载体,测序正确后采用同源重组、两次交换的方法构建突变体。采取剪叶法,在TP309水稻叶片上接种野生型菌株PXO99A 和mutL 突变体,14 d后量取从剪口处到枯斑内部边界的长度,统计病斑长度与数量。
-
PXO99A、△mutL菌株培养(PSA培养基、28 ℃、180 r·min−1)36 h后,按1∶100转接于10 mL PSA中,用全自动微生物生长曲线分析仪每2 h测定1次OD600值。
-
将PXO99A、ΔmutL菌株用PSA培养36 h,M4M培养基清洗2遍,调至OD600=1.0,再稀释至10−7共8个梯度,取2 μL菌悬液点板(PSA和M4M固体培养基)28 ℃倒置培养,2~3 d拍照。
-
PXO99A、ΔmutL菌株用PSA培养36 h后,1∶100转接于500 mL PSA中,待培养到对数生长期,ddH2O清洗1次,调OD600=1.0,取相同体积菌液,用终浓度40 mmol·L−1的H2O2处理菌液30 min,ddH2O清洗过氧化氢2遍,最后用6 mL ddH2O重悬,取出100 μL 菌液稀释到10−7、10−8后,稀释液涂PSA平板,计算菌落总数(A);其余菌液(5.9 mL)涂布于PSA加利福平的平板上,28 ℃ 培养3 d,统计菌落数(B)。B与A的比值即为突变率。
-
PCR扩增全长mutL,产物回收后无缝克隆到HindIII/EcoRI酶切的广宿主载体pHM1上。PCR验证转化大肠杆菌获得的单菌落,对阳性克隆进行测序分析。测序正确的质粒用电击法转入到ΔmutL,获得互补菌株(C-ΔmutL)。
-
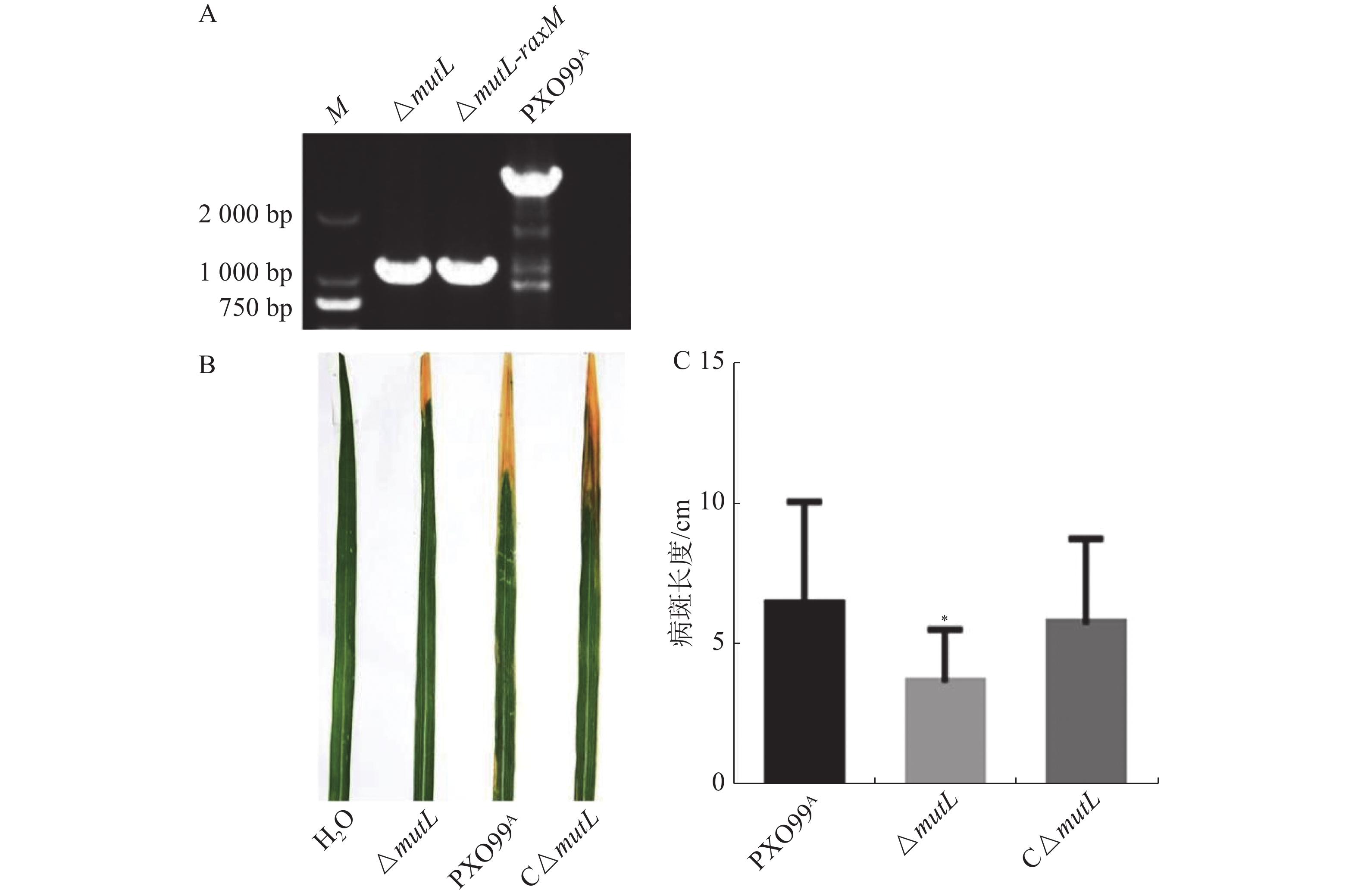
以PXO99A为野生型菌株,经过2次同源重组后mutL基因被缺失突变。以mutL两侧引物进行PCR,以PXO99A和ddH2O作对照。如图1-A所示,mutL突变体的扩增片段比野生型扩增片段短,约1 800 bp,正好符合mutL基因片段大小(1 878 bp),表明mutL突变体(ΔmutL)构建成功。为了进一步分析mutL的功能,构建了mutL的互补菌株C-ΔmutL。将PXO99A、mutL突变体及其互补菌株接种TP309水稻,结果显示mutL突变体的病斑长度比野生型及其互补菌株都短,野生型和互补菌株间没有显著差异(图1-B、C)。说明mutL正调控Xoo的致病力。
-
MutL的功能是对细胞DNA错配进行修复,维持细胞生命活动的正常进行,其突变可能改变其生长。因此,本研究检测了mutL突变是否影响Xoo的生长速度。结果显示,在丰富性培养基(PSA)和营养营养缺陷型培养基(M4M)中,ΔmutL的生长速率都略低于PXO99A(图2- A和图2-B),但没有显著差异。说明MutL在正常情况下并不显著影响Xoo的生长。
-
H2O2是常见的活性氧种类,与二价铁离子发生Fenton反应,反应产物中的羟自由基具有极强的氧化能力,会对DNA产生损伤。细胞MMR系统可以修复损伤DNA,使其遗传稳定。但突变mutL后,MMR系统受损,细胞DNA修复功能会减弱,可能增加细菌的突变率。突变率通常采用菌株利福平抗性获得率来表示[19]。因此,将对数生长期的野生型PXO99A和ΔmutL菌株进行H2O2胁迫处理,然后涂布在含利福平的平板上生长。3 d后的统计结果显示,与PXO99A相比,△mutL的突变率显著增加(图2- C),说明MutL参与Xoo的DNA修复。
-
在病原菌侵染过程中,病菌自身会改变其代谢活性、毒素合成和分泌能力等,逃避宿主攻击[18-20]。同时宿主在对抗病原菌时也会产生多种防御机制[21]。其中快速产生活性氧(ROS)限制细菌生长繁殖是植物普遍的防御机制[21]。细菌成功侵染必须要快速适应这种高ROS环境。一方面,病原菌在感染植物过程中会诱发ROS清除酶表达,进而清除ROS;另一方面,细菌会修复由ROS带来的损伤,包括DNA和其他大分子物质,帮助病原菌适应宿主环境并成功繁殖[21]。因此,DNA损伤修复对病原菌成功侵染是重要的。在细菌中,MutL/MutS系统是主要的修复系统,其突变将导致细菌自发突变率增加、抗逆性降低、环境适应能力减弱[3]。对病原菌而言,MutL/MutS可能对病原菌适应宿主体内环境起到重要的作用。
本研究结果表明,mutL突变导致Xoo在水稻TP309上的致病力降低(图1),说明MutL调控Xoo的侵染过程,是Xoo致病性相关的调节因子。同时,在正常条件下(无论是在丰富型还是营养缺陷型培养基中),mutL突变都未显著改变Xoo的生长状况(图2),说明MutL可能不是直接调控细菌生长来影响其致病力的。如前所述,在侵染过程中植物会释放大量ROS对细菌生长和DNA造成损伤[21]。本研究结果也显示,ROS增多导致mutL突变体的突变率急剧增加(图2)。这暗示MutL调控Xoo致病力可能是由于其修复DNA能力降低导致的,但具体机制还需进一步研究。
在大肠杆菌中, mutL突变后自发突变率比野生型高300倍[22]。分支杆菌mutL突变也导致自发突变率增加4倍[23]。同样,缺失mutL的耐辐射奇球菌的自发突变率也增加了16倍[24]。然而,本研究在Xoo中并未检测到野生型和mutL突变体自发突变率的差异,暗示Xoo中可能存在其他主要的DNA修复系统,而MutL是否只有在逆境环境中才发挥功能,需进一步研究证实。
MutL is required for
-
摘要: 为了了解水稻白叶枯病菌(Xanthomonas oryzae pv. oryzae,Xoo)中 的MutL/MutS系统是否参与DNA修复以及是否调控病原菌的致病力,构建了mutL突变体,比较了野生型与mutL突变体的突变率差异,野生型、mutL突变体及其互补菌株侵染水稻的能力。结果显示,mutL突变导致Xoo在H2O2胁迫下的突变率显著升高,mutL突变导致Xoo致病力下降,表明DNA错配修复蛋白MutL参与Xoo的DNA修复并正调控Xoo致病力。Abstract: Environmental stimulus and DNA replication may cause DNA mismatch and thus genomic instability, which must be repaired by organisms. All organisms evolve DNA repair systems to cope with these damages. MutL/MutS is the key DNA repair system of bacteria, but its function remains unknown in Xanthomonas oryzae pv. oryzae (Xoo). Whether MutL/MutS is involved in DNA repair and virulence in Xoo needs further studies. MutL mutant and its complementary strain were constructed, and their virulence and mutation rates were analyzed. Results showed that MutL mutation increased Xoo mutation rates under H2O2 stress, indicating that MutL is important for DNA repair during the stress. It was also found that MutL mutation reduced Xoo infection ability in rice leaves, suggesting that MutL is required for Xoo-rice interaction. In summary, MutL is involved in DNA repair in Xoo, and is also vital for Xoo virulence.
-
Key words:
- Xanthomonas oryzae pv. oryzae /
- MutL /
- virulence /
- mutation rate
-
表 1 本实验引物
引物名 序列(5′-3′) 用途 产物大小/bp PX0410ddFF GTCGAAGCTTGCCAATGTGTTGCTCGACC mutL deletion 540 PX0410ddFR TAGCGGATCCTGACGGATCGCCATCAGCGC PX0410ddRF TAGCGGATCCGGCTCGGTTTACGCTGAGCG 572 PX0410ddRR CGTGGAATTCATGCTTGACTCGCAGCGCAT PX0410hmF CAGCAAGCTTGATGGCGATCCGTCAGTTGC ΔmutL complementation 1 918 PX0410hmR GAATTCAAGCGTAGTCTGGGACGTCGTATG
GGTAACGTCCCCTGAGAAACCAAC -
[1] BRADFORD K C, WILKINS H, HAO P, et al. Dynamic human MutSalpha-MutLalpha complexes compact mismatched DNA [J]. Proc Natl Acad Sci U S A, 2020, 117(28): 16302 − 16312. doi: 10.1073/pnas.1918519117 [2] CHATTERJEE N, WALKER G C. Mechanisms of DNA damage, repair, and mutagenesis [J]. Environ Mol Mutagen, 2017, 58(5): 235 − 263. [3] CASTANEDA-GARCIA A, PRIETO A I, RODRIGUEZ-BELTRAN J, et al. A non-canonical mismatch repair pathway in prokaryotes [J]. Nat Commun, 2017, 8(14246): 14246. [4] BAN C, JUNOP M, YANG W. Transformation of MutL by ATP binding and hydrolysis: a switch in DNA mismatch repair [J]. Cell, 1999, 97(1): 85 − 97. [5] HALL M C, SHCHERBAKOVA P V, KUNKEL T A. Differential ATP binding and intrinsic ATP hydrolysis by amino-terminal domains of the yeast Mlh1 and Pms1 proteins [J]. J Biol Chem, 2002, 277(5): 3673 − 3679. [6] GROOTHUIZEN F S, SIXMA T K. The conserved molecular machinery in DNA mismatch repair enzyme structures [J]. DNA Repair (Amst), 2016, 38: 14 − 23. [7] QIU R, DEROCCO V C, HARRIS C, et al. Large conformational changes in MutS during DNA scanning, mismatch recognition and repair signalling [J]. EMBO J, 2012, 31(11): 2528 − 40. doi: 10.1038/emboj.2012.95 [8] GROOTHUIZEN F S, WINKLER I, CRISTOVAO M, et al. MutS/MutL crystal structure reveals that the MutS sliding clamp loads MutL onto DNA [J]. Elife, 2015, 4: e06744. doi: 10.7554/eLife.06744 [9] DUPES N M, WALSH B W, KLOCKO A D, et al. , Mutations in the Bacillus subtilis beta clamp that separate its roles in DNA replication from mismatch repair [J]. J Bacteriol, 2010, 192(13): 345234 − 63. [10] SIMMONS L A, DAVIES B W, GROSSMAN A D, et al. , Beta clamp directs localization of mismatch repair in Bacillus subtilis [J]. Mol Cell, 2008, 29(3): 291 − 301. doi: 10.1016/j.molcel.2007.10.036 [11] PILLON M C, LORENOWICZ J J, UCKELMANN M, et al. Structure of the endonuclease domain of MutL: unlicensed to cut [J]. Mol Cell, 2010, 39(1): 145 − 51. doi: 10.1016/j.molcel.2010.06.027 [12] PILLON M C, MILLER J H, GUARNE A. The endonuclease domain of MutL interacts with the beta sliding clamp [J]. DNA Repair (Amst), 2011, 10(1): 87 − 93. [13] SHEN M, ZHANG H, SHEN W, et al. , Pseudomonas aeruginosa MutL promotes large chromosomal deletions through non-homologous end joining to prevent bacteriophage predation [J]. Nucleic Acids Res, 2018, 46(9): 4505 − 4514. doi: 10.1093/nar/gky160 [14] 白璐, 吴利英, 吕艳红, 等. MLH1、PRMT5在卵巢癌组织中的表达及意义[J]. 生殖医学杂志, 2020, 29(10): 1356 − 1361. [15] 陈芳, 王敏, 赵声远, 等. 鲍曼不动杆菌错配修复基因mutL的序列及其系统进化关系分析[J]. 中国微生态学杂志, 2017, 29(5): 514 − 520. [16] 赵尔樱, 唐乐, 周玉洁, 等. 细菌基因组稳定性和变异性之间动态平衡的自发调控: 遗传开关的作用及其分子机制[J]. 中国科学, 2013, 43(15): 361 − 367. [17] SWING J. Reclassification of the causal agents of bacterial blight (Xanthomonas campestris pv. oryzae) and bacterial leaf streak (Xanthomonas campestris pv. oryzicola) of rice as pathovars of Xanthomonas oryzae (ex Ishiyama 1922) sp. nov. nom. rev [J]. Int J System Evol Microbiol, 1990, 40: 309 − 311. [18] SHEN Y, RONALD P. Molecular determinants of disease and resistance in interactions of Xanthomonas oryzae pv oryzae and rice [J]. Microbes and Infection, 2002, 4: 1361 − 1367. doi: 10.1016/S1286-4579(02)00004-7 [19] LI C X, LIU S Y, LIU P Y, et al. The yebN leader rNA acts as a Mn2+ sensor required for the interaction of Xanthomonas oryzae and rice [J]. Mol Plant-Microbe Ineract, 2018, 31: 932 − 939. doi: 10.1094/MPMI-02-18-0043-R [20] LI T, ZHAN Z, LIN Y, et al. Biosynthesis of amino acids in Xanthomonas oryzae pv. oryzae is essential to its pathogenicity [J]. Microorganisms, 2019, 7(12): 693. doi: 10.3390/microorganisms7120693 [21] VERMA G, MONDAL K K, KULSHRESHTHA A, et al. XopR T3SS-effector of Xanthomonas oryzae pv. oryzae suppresses cell death-mediated plant defense response during bacterial blight development in rice [J]. 3 Biotech, 2019, 9: 272. [22] LEE H, POPODI E, TANG H, et al. Rate and molecular spectrum of spontaneous mutations in the bacterium Escherichia coli as determined by whole-genome sequencing [J]. Proc Natl Acad Sci U S A, 2012, 109(41): e2774 − 2783. [23] KUCUKYILDIRIM S, LONG H, SUNG W, et al. The rate and spectrum of spontaneous mutations in mycobacterium smegmatis, a bacterium naturally devoid of the postreplicative mismatch repair pathway [J]. G3 (Bethesda), 2016, 6(7): 2157 − 2163. doi: 10.1534/g3.116.030130 [24] LONG H, KUCUKYILDIRIM S, SUNG W, et al. Background mutational features of the radiation-resistant bacterium Deinococcus radiodurans [J]. Mol Biol Evol, 2015, 32(9): 2383 − 2392. doi: 10.1093/molbev/msv119 -




 下载:
下载: