-
维氏气单胞菌(Aeromonas veronii)是一种革兰氏阴性兼性厌氧菌,隶属于气单胞菌科、气单胞菌属[1]。有研究表明,这种新兴的水生病原菌可以引起人类和动物(包括水生动物)的腹泻、伤口感染和出血性败血症[2-3]。它兼具肠毒素、Ⅲ型分泌效应蛋白AexU、组氨酸激酶BvgS、丝氨酸蛋白酶、外膜蛋白和鞭毛等多种毒力因子[4-5],能给水产养殖业造成巨大的经济损失,并对食品安全造成威胁[6-7]。例如:养殖场的鲶鱼和黄颡鱼感染维氏气单胞菌后,会大量死亡[8-9]。因此,维氏气单胞菌引起的鱼类细菌性疾病是制约水产养殖发展的重要因素,而解构维氏气单胞菌的抗逆性对维氏气单胞菌的疾病防治具有重要意义。类CysB蛋白Cbl(CysB like protein)是LysR家族的转录调控因子。1995年,首次在大肠杆菌中发现了cbl基因,它与cys调控子的转录调控因子CysB蛋白非常相似,与CysB具有40%的氨基酸序列同源性[10]。在硫代谢中,Cbl作为细胞内硫酸盐水平的传感器,在体内和体外激活tau和ssu启动子,控制牛磺酸和脂肪族磺酸盐的转运及脱硫所需的tauABCD和ssuEADCB的转录,从而稳定细菌体内的硫酸盐水平[11-13]。Cbl的功能与CysB高度同一,目前对Cbl的研究较少,但鉴于CysB现有的功能,可以推断Cbl存在的潜在毒力控制、参与细菌体内氧化应激机制等重要功能[14-16]。因此,本研究以维氏气单胞菌为研究对象,对Cbl蛋白进行原核表达和纯化,旨在为进一步揭示Cbl蛋白的功能和机制奠定基础。
-
维氏气单胞菌(Aeromonas veronii)C4菌株、大肠杆菌BL21(DE3)以及含pET-28a质粒的大肠杆菌DH5α均为课题组保存菌株。
-
Bacteria DNA Extraction Kit、FastPure Gel DNA Extraction Mini Kit、FastPure Plasmid Mini Kit、2 × Phanta Max Master Mix以及2 × Rapid Taq Master Mix均购自Vazyme公司;限制性内切酶(EcoRI-HF和HindIII-HF)及Fast T4 DNA Ligase /Buffers均购自NEB公司;卡那霉素(Kan,水溶,储存的质量浓度为50 g·L−1,工作质量浓度为50 mg·L−1,−20 ℃保存)、异丙基−β−D 硫代半乳糖苷(IPTG,水溶,储存浓度为0.1 mol·L−1,工作浓度为0.1 mmol·L−1,−20 ℃保存)、聚丙烯酰胺混合液(29∶1,体积比)、Tris、乙二胺四乙酸(EDTA)、十二烷基硫酸钠(SDS)、过硫酸铵(APS)、四甲基乙二胺(TEMED)以及LB培养基(w=1%NaCl、w=1%蛋白胨和w=0.5%酵母提取物)均购自Solarbio公司。
Eppendorf Centrifuge 5418高速台式离心机(Eppendorf 中国有限公司)、Life ECO-PCR 基因扩增仪(杭州博日科技有限公司)、Bio-Rad MicroPulser电穿孔仪(美国Bio-Rad 公司)、HZQ-F100 振荡培养箱(哈尔滨市东联生化仪器有限公司)、SYNERGY-H1多功能酶标仪(美国BioTek 公司)、PowerPac™电泳仪(美国Bio-rad 公司)以及Typhoon FLA 9500多功能生物分子成像仪(浙江德力西电器有限公司)。
-
设计cbl基因扩增引物(表1)。使用Bacteria DNA Extraction Kit提取维氏气单胞菌C4总DNA,以维氏气单胞菌C4基因组DNA为模板,利用cbl-F和cbl-R对cbl基因片段进行PCR扩增,其片段大小为948 bp。PCR反应体系为2 × Phanta Max Master Mix 25 μL,cbl-F(10 μmol·L−1)1 μL,cbl-R(10 μmol·L−1)1 μL,基因组DNA 1 μL,ddH2O 22 μL,反应总体积为50 μL。反应程序:94 ℃预变性10 min,94 oC变性30 s,55 ℃退火30 s,72 ℃延伸1 min,共30个循环,72 ℃后延伸10 min。PCR完成后进行琼脂糖凝胶(w=0.5%)电泳鉴定,再用FastPure Gel DNA Extraction Mini Kit纯化回收扩增的DNA片段。
表 1 引物序列
Table 1. Primer sequence
引物 Primers 核苷酸序列(5′→3′) Nucleotide sequence (5′→3′) 长度/bp Length cbl-F CCGGAATTCGTGAATTTTCAGCAGTTACG 29 cbl-R CCCAAGCTTTCAAATCTGGAAGTCGAG 27 -
使用FastPure Plasmid Mini Kit提取pET-28a质粒,再用EcoRI-HF和HindⅢ-HF对纯化回收的DNA片段和pET-28a质粒进行双酶切,酶切产物使用FastPure Gel DNA Extraction Mini Kit纯化回收后按一定比例(质粒∶片段的摩尔比为1∶5)使用Fast T4 DNA Ligase过夜连接,将扩增出的cbl基因连入pET-28a载体的EcoRI(GAATTC)~ HindⅢ(AAGCTT)位点之间。将连接产物通过电击转入大肠杆菌BL21(DE3)感受态细胞中,复苏后涂布于含卡那霉素LB平板,过夜培养后挑菌验证,阳性克隆子送往生工生物工程(上海)股份有限公司测序。
-
将含有pET-28a-Cbl重组载体的大肠杆菌BL21(DE3)菌株接种至含有50 mg·L−1卡那霉素的LB液体培养基中,在37 ℃摇床培养至OD600在0.4~0.6之间,加入IPTG至终浓度为100 μmol·L−1,在150 r·min−1、15 ℃条件下,过夜诱导表达12 h。然后将菌液冷冻离心(4 ℃,4 000 r·min−1)10 min,用10 mL的 Tris-HCl(50 mmol·L−1 Tris,350 mmol·L−1 NaCl)缓冲液重悬菌体。超声破碎仪设置:超声时间5 s,间隙时间7 s,总时间25 min,温度25~28 ℃,功率35%。冰上超声破碎重悬的菌体后将破碎的菌体冷冻离心(4 ℃,12 000 r·min−1)10~15 min,分离得到沉淀和上清。提前取出镍柱,用20 mL Tris-HCl缓冲液清洗2~3遍后关闭镍柱阀门,将破碎上清注入镍柱中并封口,置于冰上150 r·min−1孵育1 h。打开镍柱阀门,流尽孵育上清,用50 mmol·L−1咪唑洗脱杂蛋白,100 mmol·L−1咪唑收集目的Cbl蛋白,500 mmol·L−1咪唑清洗镍柱。最后进行SDS聚丙烯酰胺凝胶电泳(SDS-PAGE)分析。
-
剪下合适大小的透析袋,放入含有1 mmol·L−1 EDTA-2Na的w=2% NaHCO3溶液中煮10 min,去离子水清洗后放入1 mmol·L−1 EDTA-2Na溶液中再次煮10 min,再次用去离子水清洗透析袋。将诱导表达的蛋白加入透析袋中,用含有φ=30%甘油的Tris-HCl作为透析液于4 ℃进行透析,每6 h更换1次透析液,共透析12 h。取出透析袋中透析好的蛋白,SDS-PAGE观察目的条带,用BOSTER的BCA蛋白浓度测定试剂盒测量Cbl蛋白质量浓度。
-
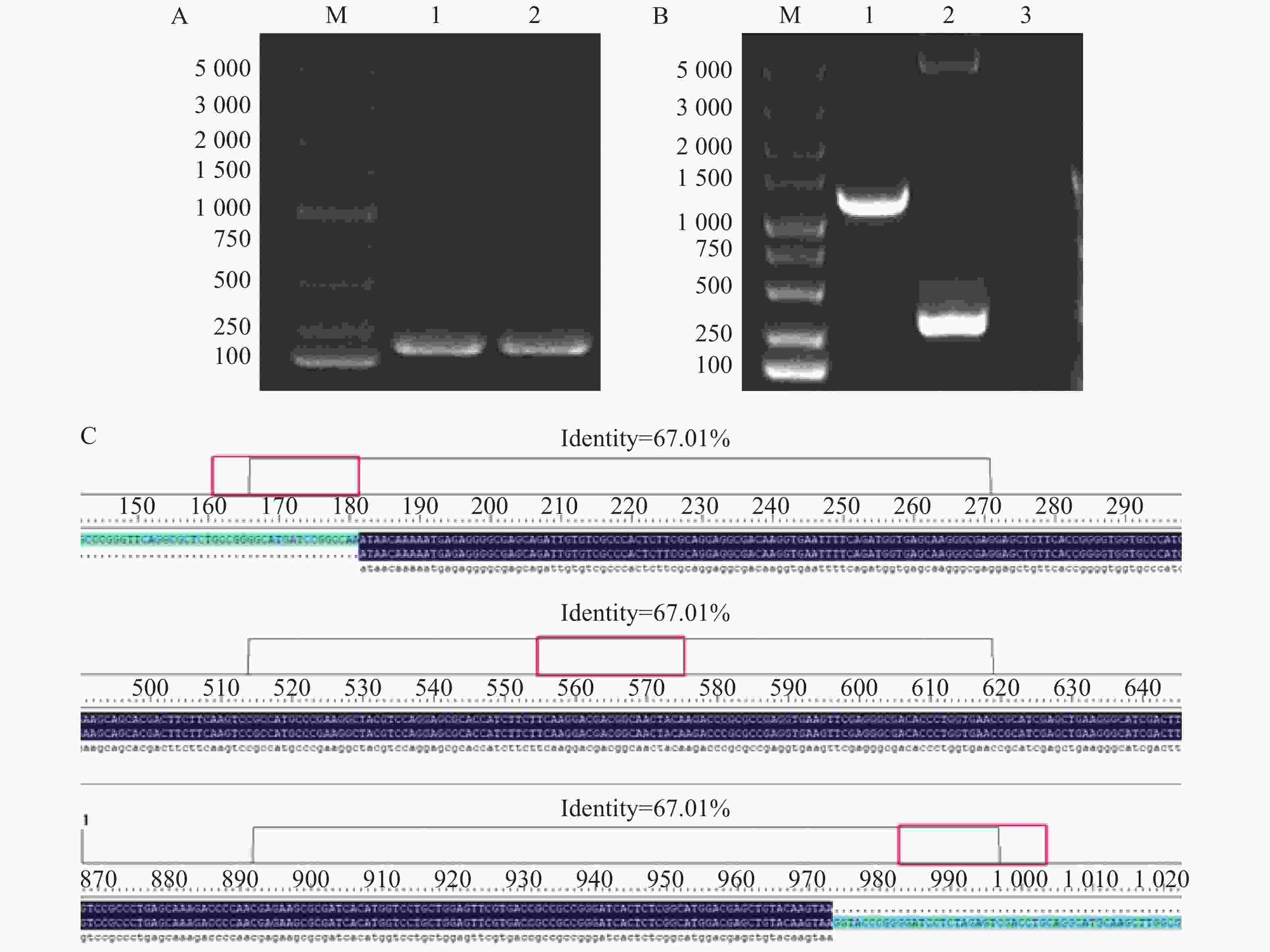
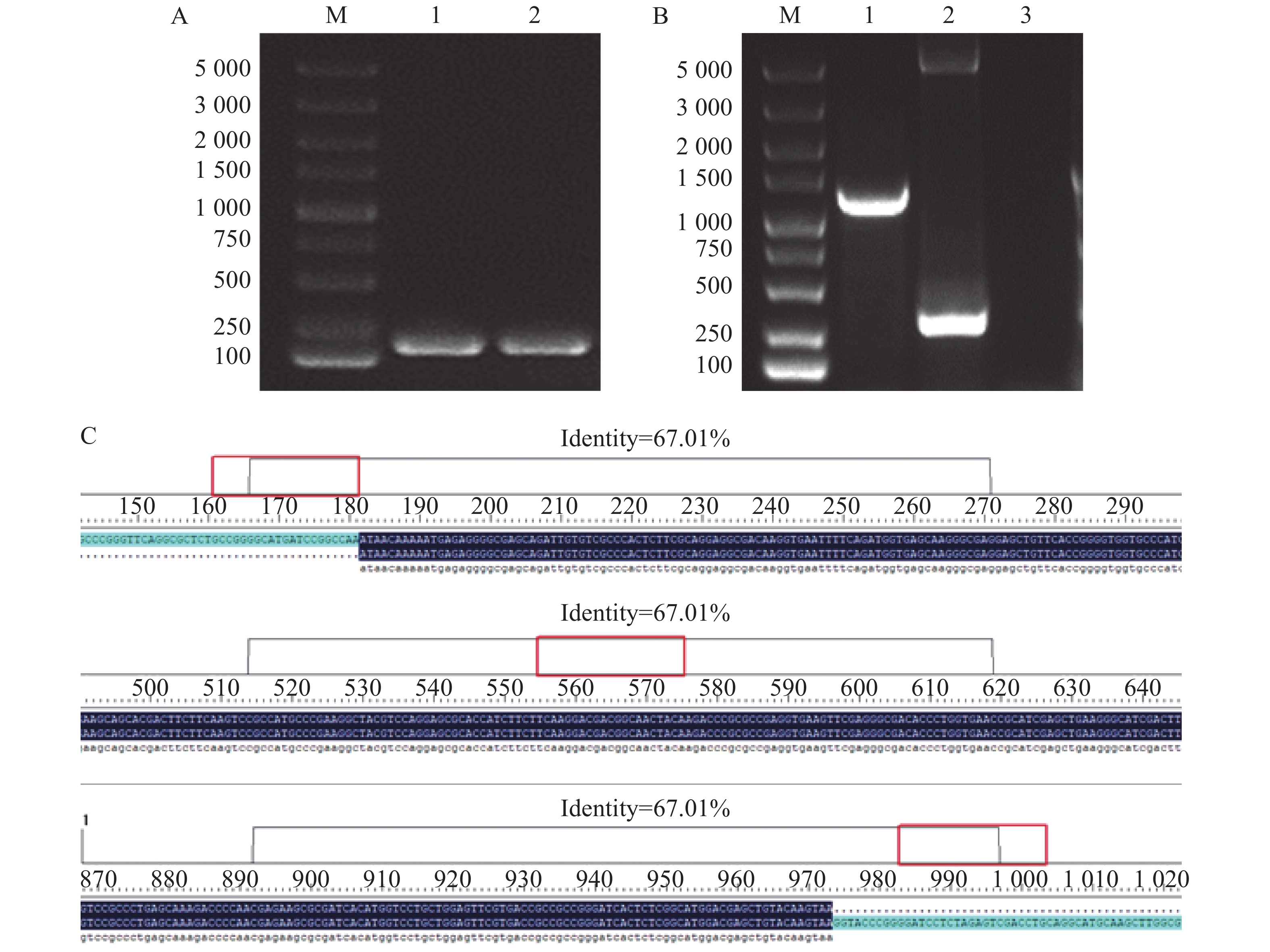
使用Bacteria DNA Extraction Kit提取维氏气单胞菌C4基因组DNA,并用维氏气单胞菌特异性引物进行PCR验证,维氏气单胞菌C4破碎菌体作为阳性对照。从图1-A可知,提取的维氏气单胞菌C4基因组DNA条带单一清晰无杂带,大小与阳性对照相比一致,约为200 bp,符合理论预期。而后经功能酶标仪检测维氏气单胞菌C4基因组DNA浓度为672.76 μg·L−1,可用于后续实验。
将双酶切后的维氏气单胞菌cbl基因扩增片段和pET-28a质粒双酶切片段用Fast T4 DNA Ligase酶连,电击转化法转入大肠杆菌BL21(DE3)感受态细胞中并涂布于含有50 mg·L−1卡那霉素的LB平板上进行筛选培养,对平板上长出的单菌落使用pET-28a载体引物进行PCR验证。从图1-B可知,挑取的阳性克隆子的条带处于1 000~1 500 bp之间,条带单一清晰无杂带,且与以空载的pET-28a质粒作为模板的阳性对照相比相差约948 bp,符合理论预期,说明阳性克隆子成功的将cbl基因片段连入pET-28a载体。
富集培养后提取质粒送往生工生物工程(上海)股份有限公司测序。测序结果用DNAMAN进行分析,结果如图1-C所示,阳性克隆子序列与维氏气单胞菌C4基因组中cbl序列一致。进一步验证了成功构建原核重组表达质粒pET-28a-Cbl。
-
将含有重组pET-28a-Cbl载体的大肠杆菌BL21(DE3)于37 ℃培养至OD600为0.4~0.6后,加入IPTG于37 ℃诱导表达2、4、6 h,以未加IPTG诱导的pET-28a-Cbl以及加入IPTG的空载pET-28a作为对照,对得到的样品进行SDS-PAGE凝胶电泳分析Cbl蛋白在上清和沉淀中的表达情况。从图2可见,于37 ℃条件下,经IPTG诱导后,pET-28a-Cbl在35.35 kDa左右有1条明显的特异性的蛋白条带,并且条带浓度明显大于对照组,且2、4、6 h这3个时间点的蛋白诱导量随时间的增长而变大,但是Cbl蛋白结构不稳定,易出现错配的二硫键,导致包涵体的形成,因此,Cbl蛋白大多在沉淀中表达,在上清中只有少量表达。为了解决该问题,在相同条件下加入IPTG后,15 ℃低温诱导表达,以Tris-HCl作为缓冲液超声破碎细胞,破碎得到的上清蛋白使用镍柱纯化。使用不同浓度的咪唑(50、100、150、200、300、400、500 mmol·L−1)进行梯度洗脱,对得到的样品进行SDS-PAGE凝胶电泳分析。分析结果(图3)表明:经过低温诱导后,Cbl蛋白在上清中表达量明显增加,镍柱孵育后大小为35.35 kDa,带有His标签的Cbl蛋白成功与镍柱结合,并且比较不同浓度咪唑洗脱得到的蛋白,其中,100 mmol·L−1咪唑洗脱时能将大部分目的蛋白洗脱下来,此时蛋白条带单一浓度较高,能用于下一步实验。
-
使用含φ=30%甘油的Tris-HCl缓冲液作为透析液,4 ℃过夜透析Cbl蛋白充分去除咪唑。取出透析蛋白,用BOSTER的BCA蛋白浓度测定试剂盒制作标准曲线,测量Cbl蛋白OD562值,进而根据标准曲线得到Cbl蛋白的浓度。制作的标准曲线如图4所示,横坐标表示蛋白的OD562,纵坐标表示蛋白质量浓度(μg·L−1,标准曲线R2值=0.9676,表明标准曲线可信度高,透析所得蛋白OD562=0.555,根据公式y=1 112x−237.74,代入计算可得Cbl蛋白质量浓度为601.405 mg·L−1。
-
渔业在我国国民经济贸易中的作用日益增强,水产养殖业已经成为促进渔业可持续、快速、稳定发展的重要力量[17]。然而,鱼类细菌性疾病的频繁发生是制约水产养殖发展的重要因素[7]。近年来,维氏气单胞菌大规模爆发的病例越来越多。有报道称,维氏气单胞菌可感染淡水鱼[18-19]、两栖动物[20]和哺乳动物[2],给养殖业造成严重的经济损失,并威胁食品安全。此外,人们也可能通过接触维氏气单胞菌污染的表面而感染,如被污染的家畜和家禽肉类和海产品,从而导致肠胃炎[21]。但是维氏气单胞菌致病机制复杂,其致病机理尚未研究透彻,治疗维氏气单胞菌引起的各种疾病显得至关重要。
Cbl作为硫酸盐水平的感应器,在硫代谢途径中有着重要作用。在大肠杆菌中,迄今为止鉴定的ssi基因包括ssuEADCB和tauABCD操纵子,分别编码有机硫化合物、脂肪族磺酸盐和牛磺酸摄取系统的,以及从各自底物中释放亚硫酸盐的氧化还原酶型酶[22]。tau和ssu的表达仅在无无机硫酸盐的情况下诱导,且需要转录因子Cbl的存在[23]。尽管与CysB高度同源,但与CysB不同的是Cbl不需要诱导配体就能从目标启动子中激活转录,其功能受到硫酸同化途径中的第一个中间体腺苷5−磷酸硫酸酯(APS)的负调控[24]。因此,cbl介导的调节解释了大肠杆菌利用硫源的层次:半胱氨酸>硫酸盐>磺酸盐[25]。本实验室的前期研究表明,Cbl与维氏气单胞菌H2S的合成密不可分,而在细菌中,H2S参与了细菌对活性氧(ROS和抗生素诱导的氧化损伤的防御[26]。因此,Cbl对维氏气单胞菌的氧化耐受也起着重要作用。
本研究成功构建了Cbl蛋白编码基因cbl的原核表达载体pET-28a-Cbl,并根据Cbl与CysB蛋白的高同源性,参考CysB的蛋白表达条件[17],摸索出了Cbl蛋白诱导表达和纯化的适宜条件。本研究表明,Cbl蛋白在37 ℃时,主要在细菌裂解后沉淀中表达,易形成包涵体,所以对Cbl蛋白进行低温诱导,减少包涵体的形成。因此,得出Cbl蛋白诱导的适宜条件:诱导时间为8 h,IPTG终浓度为0.1 mmol·L−1,诱导温度为15 ℃。Cbl蛋白经由Ni柱纯化后主要分布于100 mmol·L−1咪唑的洗脱液中,纯化得到的Cbl蛋白条带清晰单一,透析后蛋白浓度较高,能满足后续功能研究实验的要求。
本研究对Cbl进行了原核表达载体的构建以及蛋白表达纯化,最后得到理想的Cbl蛋白,为下一步进行Cbl蛋白的功能及其分子机制研究奠定了基础,也为维氏气单胞菌氧化耐受抗逆性的研究提供了理论依据。
Prokaryotic Expression and Purification of Cbl Protein from Aeromonas veronii
-
摘要: 维氏气单胞菌(Aeromonas veronii)引起的细菌性疾病已威胁到鱼类养殖业的发展。Cbl(类CysB蛋白)是细菌硫代谢的重要调控因子之一。本研究选取pET-28a为Cbl的原核表达载体,以大肠杆菌BL21(DE3)为宿主菌,使用IPTG(Isopropyl-beta-D-thiogalactoside,异丙基−β−D−硫代半乳糖苷)诱导表达,用SDS-PAGE凝胶电泳验证蛋白条带,再用BOSTER的BCA蛋白浓度测定试剂盒测量Cbl蛋白浓度,寻找较适宜的蛋白表达条件。结果表明,当诱导时间为8 h,IPTG终浓度为0.1 mmol·L−1,诱导温度为15 ℃,咪唑洗脱浓度为100 mmol·L−1时,Cbl蛋白质量浓度能达到601.405 mg·L−1,可满足后续实验要求。Abstract: Bacterial diseases caused by Aeromonas veronii have threatened the development of fish production. And Cbl (CysB like protein) is one of the important regulatory factors of bacterial sulfur metabolism. Escherichia coli BL21 (DE3) was selected as the prokaryotic expression vector for Cbl, and culturedadded with IPTG (Isopropyl-beta-D-thiogalactoside) for culture to induce the expression of Cb1. The protein samples were run on SDS-PAGE gel electrophoresis to verify the protein bands of Cb1, and then the concentration of Cbl was determined with Boster’s BCA protein assay kit. The results showed that Cbl was successfully purified, and that the concentration of Cbl was up to 601.405 mg·L−1, which meets the requirements for subsequent experiments, when the induction time was 8h, the final IPTG concentration was 0.1 mmol·L−1, the induction temperature was 15 ℃ and the Imidazole concentration in the elution buffer was 100 mmol·L−1.
-
Key words:
- Cbl /
- vector construction /
- prokaryotic expression /
- protein purification
-
图 2 不同条件下Cbl蛋白的表达图谱
M:低分子质量蛋白Marker(14.4~97.4 KDa);1:诱导后培养2 h的沉淀;2:诱导培养后2 h的上清;3:诱导后培养4 h的沉淀;4:诱导后培养4 h的上清;5:诱导后培养6 h的沉淀;6:诱导后培养6 h的上清;7:空载pET-28a诱导后培养2 h的沉淀;8:未加IPTG诱导培养2 h的沉淀。
Fig. 2 Expression profile of Cbl protein under different conditions
M: Protein Marker (14.4kD-97.4 kDa); 1: The precipitation of the induced sample cultured for 2 h; 2: The supernatant of the sample cultured for induction for 2 h; 3: The precipitation of the sample cultured for induction for 4 h; 4: The supernatant of the sample cultured for induction for 4 h; 5: The precipitation of the sample cultured for induction for 6 h; 6: The supernatant of the sample cultured for induction for 6 h; 7: The precipitation of the sample containing pET-28a empty vector cultured for induction for 2 h; 8: The precipitation of the sample cultured for induction for 2 h without IPTG
图 3 Cbl蛋白的纯化图谱
M:低分子质量蛋白Marker(14.4~97.4 kDa);1:超声破碎后的上清;2:镍柱孵育后的过柱蛋白;3:50 mM咪唑洗脱蛋白;4:100 mmol·L−1咪唑洗脱蛋白;5:150 mmol·L−1咪唑洗脱蛋白;6:200 mmol·L−1咪唑洗脱蛋白;7:300 mmol·L−1咪唑洗脱蛋白;8:400 mmol·L−1咪唑洗脱蛋白;9:500 mmol·L−1咪唑洗脱蛋白。
Fig. 3 Purification map of Cbl protein
M: Protein Marker (14.4-97.4 kDa); 1: Supernatant after ultrasonic crushing; 2: Over-column proteins after Ni column incubation; 3: Protein eluted with 50 mmol·L−1 imidazole; 4: Protein eluted with 100 mmol·L−1 imidazole; 5: Protein eluted with 150 mmol·L−1 imidazole; 6: Protein eluted with 200 mmol·L−1 imidazole; 7: Protein eluted with 300 mmol·L−1 imidazole; 8: Protein eluted with 400 mmol·L−1 imidazole; 9: Protein eluted with 500 mmol·L−1 imidazole.
表 1 引物序列
Table 1 Primer sequence
引物 Primers 核苷酸序列(5′→3′) Nucleotide sequence (5′→3′) 长度/bp Length cbl-F CCGGAATTCGTGAATTTTCAGCAGTTACG 29 cbl-R CCCAAGCTTTCAAATCTGGAAGTCGAG 27 -
[1] 曹欣, 唐燕琼, 李宏, 等. Flag-ALD融合载体构建及蛋白表达[J]. 热带生物学报, 2020, 11(2): 132 − 137. [2] BHOWMICK U D, BHATTACHARJEE S. Bacteriological, Clinical and virulence aspects of Aeromonas-associated diseases in humans [J]. Polish Journal of Microbiology, 2018, 67(2): 137 − 149. doi: 10.21307/pjm-2018-020 [3] LI T, WANG Z, HAN H, et al. Dual Antibacterial activities and biofilm eradication of a marine peptide-N6NH2 and its analogs against multidrug-resistant Aeromonas veronii [J]. International Journal of Molecular Sciences, 2020, 21(24): 1 − 28. [4] SONG M F, KANG Y H, ZHANG D X, et al. Immunogenicity of extracellular products from an inactivated vaccine against Aeromonas veronii TH0426 in Koi, Cyprinus Carpio [J]. Fish and Shellfish Immunology, 2018, 81: 176 − 181. doi: 10.1016/j.fsi.2018.07.004 [5] REYES-BECERRIL M, ANGULO C, ASCENCIO F. Humoral immune response and TLR9 gene expression in Pacific red snapper (Lutjanus peru) experimentally exposed to Aeromonas veronii [J]. Fish and Shellfish Immunology, 2015, 42(2): 289 − 296. doi: 10.1016/j.fsi.2014.11.002 [6] HOAI T D, TRANG T T, VAN TUYEN N, et al. Aeromonas veronii caused disease and mortality in channel catfish in Vietnam [J]. Aquaculture, 2019, 513: 734425. doi: 10.1016/j.aquaculture.2019.734425 [7] HUANG H, ZHOU P, CHEN P, et al. Alteration of the gut microbiome and immune factors of grass carp infected with Aeromonas veronii and screening of an antagonistic bacterial strain (Streptomyces flavotricini) [J]. Microbial Pathogenesis, 2020, 143: 104092. doi: 10.1016/j.micpath.2020.104092 [8] SILVER A C, WILLIAMS D, FAUCHER J, et al. Complex evolutionary history of the Aeromonas veronii Group Revealed by Host Interaction and DNA Sequence Data [J]. PLoS ONE, 2011, 6(2): e16751. doi: 10.1371/journal.pone.0016751 [9] MOLINARI L M, DE OLIVEIRA SCOARIS D, PEDROSO R B, et al. Bacterial Microflora in the gastrointestinal tract of Nile tilapia, Oreochromis niloticus, cultured in a semi-intensive system [J]. Acta Scientiarum - Biological Sciences, 2003, 25(2): 267 − 271. [10] IWANICKA-NOWICKA R, HRYNIEWICZ M. A new gene, Cbl, encoding a member of the LysR family of transcriptional regulators belongs to Escherichia coli Cys Regulon [J]. Gene, 1995, 166(1): 11 − 17. doi: 10.1016/0378-1119(95)00606-8 [11] VAN DER PLOEG J R, IWANICKA-NOWICKA R, BYKOWSKI T, et al. The Escherichia coli ssuEADCB gene cluster is required for the utilization of sulfur from aliphatic sulfonates and is regulated by the transcriptional activator Cbl [J]. Journal of Biological Chemistry, 1999, 274(41): 29358 − 29365. doi: 10.1074/jbc.274.41.29358 [12] VAN DER PLOEG J R, IWANICKA-NOWICKA R, KERTESZ M A, et al. Involvement of CysB and Cbl regulatory proteins in expression of the TauABCD operon and other sulfate starvation-inducible genes in Escherichia coli [J]. Journal of Bacteriology, 1997, 179(24): 7671 − 7678. doi: 10.1128/JB.179.24.7671-7678.1997 [13] VAN DER PLOEG J R, EICHHORN E, Leisinger T. Sulfonate-sulfur metabolism and its regulation in Escherichia coli [J]. Archives of Microbiology, 2001, 176(1/2): 1 − 8. [14] IWANICKA-NOWICKA R, ZIELAK A, COOK A M, et al. Regulation of sulfur assimilation pathways in Burkholderia cenocepacia: Identification of transcription factors CysB and SsuR and their role in control of target genes [J]. Journal of Bacteriology, 2007, 189(5): 1675 − 1688. doi: 10.1128/JB.00592-06 [15] ÁLVAREZ R, NEUMANN G, FRÁVEGA J, et al. CysB-dependent upregulation of the Salmonella typhimurium CysJIH operon in response to antimicrobial compounds that induce oxidative stress [J]. Biochemical and Biophysical Research Communications, 2015, 458(1): 46 − 51. doi: 10.1016/j.bbrc.2015.01.058 [16] MIRONOV A, SEREGINA T, NAGORNYKH M, et al. Mechanism of H2S-mediated protection against oxidative stress in Escherichia coli [J]. Proceedings of the National Academy of Sciences of the United States of America, 2017, 114(23): 6022 − 6027. doi: 10.1073/pnas.1703576114 [17] GUI J F, ZHU Z Y. Molecular basis and genetic improvement of economically important traits in Aquaculture Animals [J]. Chinese Science Bulletin., 2012, 57(15): 1751 − 1760. doi: 10.1007/s11434-012-5213-0 [18] LI T, YANG B, SUN Y, et al. Aeromonas veronii infection in commercial freshwater fish: A potential threat to public health [J]. Animals, 2020, 10(4): 608. doi: 10.3390/ani10040608 [19] GONZÁLEZ-SERRANO C J, SANTOS J A, GARCÍA-LÓPEZ M L, et al. Virulence markers in Aeromonas hydrophila and Aeromonas veronii biovar sobria isolates from freshwater fish and from a diarrhoea case [J]. Journal of Applied Microbiology, 2002, 93(3): 414 − 419. doi: 10.1046/j.1365-2672.2002.01705.x [20] PEARSON M D, HIRONO I, AOKI T, et al. Virulence properties of motile Aeromonas isolated from farmed frogs Rana tigerina and R. rugulosa [J]. Diseases of Aquatic Organisms, 2000, 40(3): 185 − 193. [21] PUND R P, THEEGARTEN D. The Importance of Aeromonas as a Human Pathogen [J]. Bundesgesundheitsblatt - Gesundheitsforschung - Gesundheitsschutz, 2008, 51(5): 569 − 576. doi: 10.1007/s00103-008-0531-8 [22] VAN DER PLOEG J R, WEISS M A, SALLER E, et al. Identification of sulfate starvation-regulated genes in Escherichia coli: A gene cluster involved in the utilization of taurine as a sulfur source [J]. Journal of Bacteriology, 1996, 178(18): 5438 − 5446. doi: 10.1128/JB.178.18.5438-5446.1996 [23] MENG X, WOLFE S A. Identifying DNA sequences recognized by a transcription factor using a bacterial one-hybrid system [J]. Nature Protocols, 2006, 1(1): 30 − 45. doi: 10.1038/nprot.2006.6 [24] BYKOWSKI T, VAN DER PLOEG J R, IWANICKA-NOWICKA R, et al. The switch from inorganic to organic sulphur assimilation in Escherichia coli: Adenosine 5′-Phosphosulphate (APS) as a signaling molecule for sulphate excess [J]. Molecular Microbiology, 2002, 43(5): 1347 − 1358. doi: 10.1046/j.1365-2958.2002.02846.x [25] STEC E, WITKOWSKA-ZIMNY M, HRYNIEWICZ M M, et al. Structural basis of the sulphate starvation response in E. coli: Crystal structure and mutational analysis of the cofactor-binding domain of the Cbl transcriptional regulator [J]. Journal of Molecular Biology, 2006, 364(3): 309 − 322. doi: 10.1016/j.jmb.2006.06.033 [26] SHATALIN K, SHATALINA E, MIRONOV A, et al. H2S: A universal defense against antibiotics in bacteria [J]. Science, 2011, 334(6058): 986 − 990. doi: 10.1126/science.1209855 -
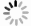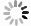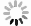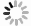



 下载:
下载: