-
作为生物体基本的结构与功能单位,细胞具有严格的代谢体系,以适应不同的环境条件、维持机体稳态与能量平衡。细胞自噬(autophagy)是一种高度保守的分解代谢途径,通过降解不必要的、不需要的或功能失调的细胞成分,包括半衰期较长的或错误折叠的蛋白质,甚至是损伤的细胞器,如线粒体、核糖体、内质网等,严密调控细胞稳态[1]。通常情况下,自噬通路的激活是由于细胞处于营养缺乏的状态,自噬体(autophagosome)包裹细胞质组分,通过分解代谢为细胞提供营养物质,此时自噬是没有选择性的,这种非选择性的自噬也被认为是经典的自噬途径(canonical autophagy)。近年来,许多研究表明,细胞倾向于有选择地借助自噬降解底物,胞内外的各种刺激因素,如细胞器受损、有毒蛋白质的聚集或氧化应激,均可诱导细胞自噬[2]。根据包裹底物的不同,具有底物选择性的自噬又被分为线粒体自噬(mitophagy)、聚集体自噬(aggrephagy)、过氧化物酶体自噬(pexophagy)、病原体自噬(xenophagy)等。病原体自噬,即细菌等病原微生物入侵时,自噬体包裹入侵的病原后与溶酶体融合,清除胞内细菌[3]。同时,根据自噬体的形成过程及运送方式的不同,可将细胞自噬分为3类:①巨自噬(macroautophagy):形成具有双层膜结构的自噬体包裹胞内物质。一般情况下所说的自噬指的是巨自噬[4]。②微自噬(microautophagy):溶酶体或液泡表面的形变直接吞没胞浆成分[5]。③分子伴侣介导的自噬(chaperone-mediated autophagy, CMA):具有KEFRQ样基序的蛋白在分子伴侣(如HSP70)的帮助下,通过LAMP-2A转运体直接转运到溶酶体[6]。
自噬在清除胞内废物、维持细胞能量水平、应对环境变化等方面都发挥着重要的作用。细菌感染细胞时,通过多种途径激活病原体自噬。细胞接受到自噬诱导信号后,在胞浆中形成多个自噬体膜发生中心,形成杯状的吞噬泡(phagophore),随后吞噬泡的双层膜结构延伸,包裹胞浆内的目标组分,封闭后形成300~900 nm的自噬体,微管相关蛋白轻链3(microtubule-associated proteins 1A and 1B, MAP1LC3/LC3)由胞浆(LC3-I)转位到自噬体膜(LC3-II)。目前认为,自噬体膜并不直接来源于高尔基体或内质网,而是在包浆中重新生成的,具体机制尚不清楚。成熟的自噬体与溶酶体(lysosome)融合形成自噬溶酶体(autolysosome),在自噬溶酶体中,内容物被降解、回收。病原体自噬激活后,一方面通过对包裹细菌的降解限制胞内细菌的增殖,另一方面,包裹细菌的自噬体也为细菌提供了便宜的生存环境,许多细菌进化出在自噬体中增殖后通过破坏自噬体膜逃逸细胞自噬的机制。本文将从细菌激活自噬途径、自噬反应通路、细菌逃逸或利用细胞自噬机制以及利用自噬治疗感染性疾病方法5个方面展开,深入阐明细菌感染与细胞自噬的动态平衡机制,以期为后续的细胞自噬研究提供资料与参考。
-
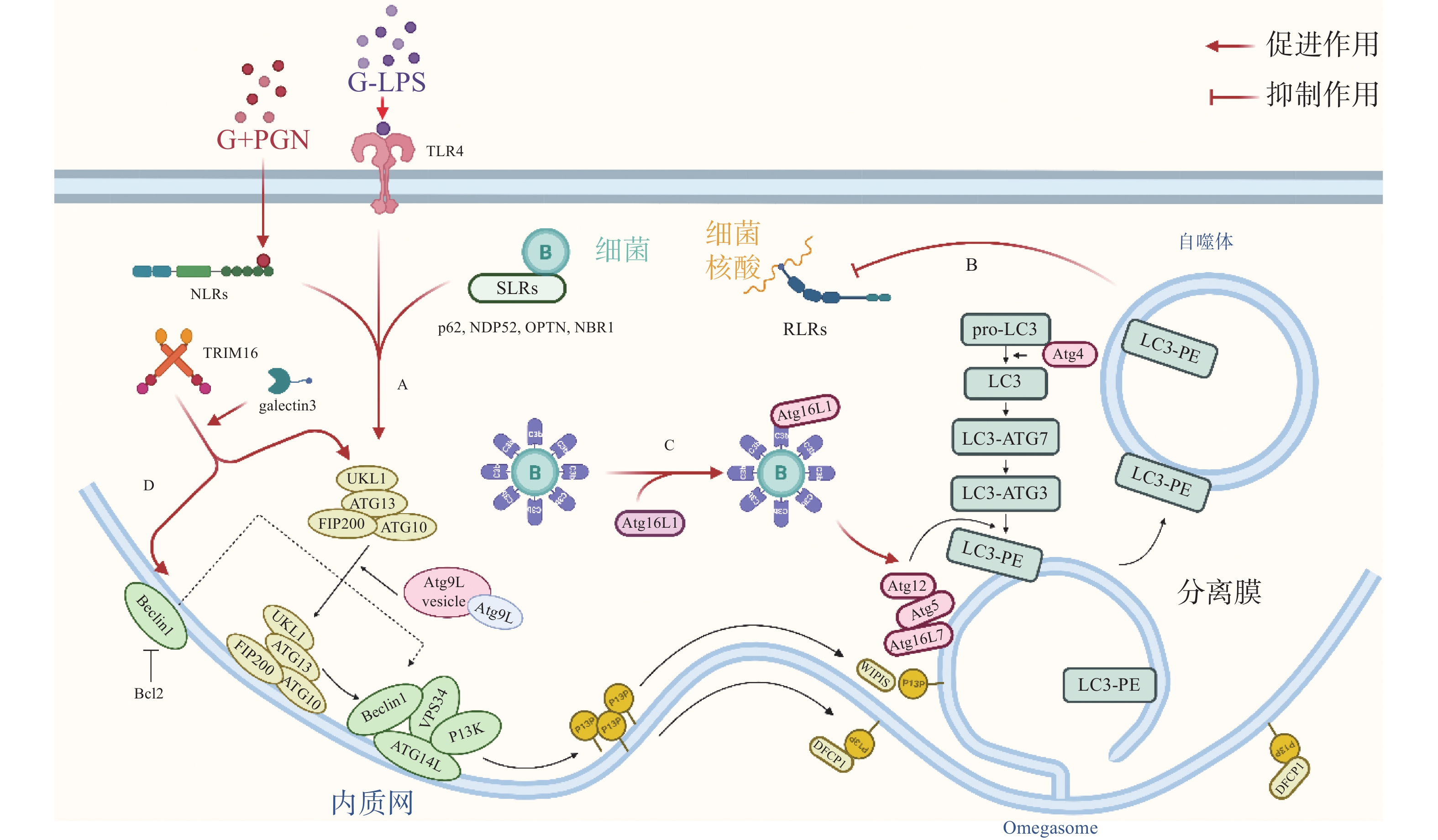
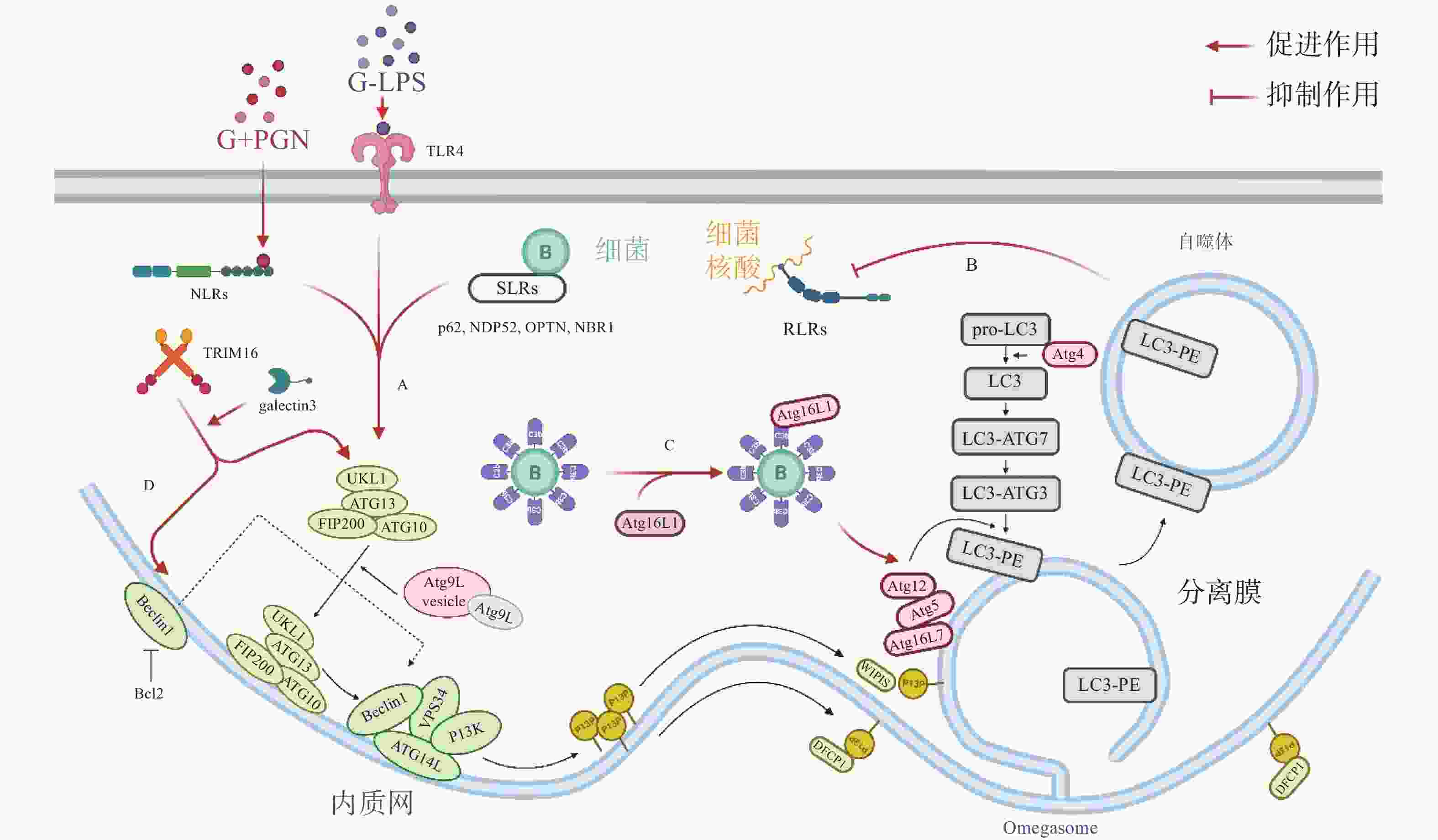
自噬体识别胞内细菌并限制其增殖,可视为一种天然免疫(innate immunity)通路。如图1-A,革兰氏阴菌外膜的脂多糖(lipopolysaccharide, LPS)或革兰氏阳菌外壁的肽聚糖(peptidoglycan, PGN)成分分别被toll样受体4(toll-like receptor 4, TLR4)或核苷酸结合寡聚结构域1(Nucleotide Binding Oligomerization Domain Containing 1, NOD1)和NOD2识别,TLRs与NOD样受体(NOD-like receptors, NLRs)激活后在细菌入侵细胞、细胞摄取或吞噬细菌时均可促进自噬[7]。如图1-B,与TLRs和NLRs不同,维甲酸诱导基因 Ⅰ 样受体(Retinoic-acid-inducible gene I like receptors,RLRs)识别病原体核酸,通过激活干扰素调节因子3(interferon regulatory factor 3, IRF)-干扰素(Interferon,IFN)激活天然免疫系统对抗病原感染,自噬负调控RLRs,清除过量产生的IFN防止长时间激活天然免疫对宿主细胞产生伤害[8]。细菌进入胞质后,还可被选择性自噬接头蛋白(sequestosome 1, SQSTM1/p62-like receptors, SLRs)识别[9],已有研究发现多种SLRs激活自噬通路,如SQSTM1 / p62、NDP52和OPTN蛋白等介导细菌识别[10],此外,参与过氧化物酶体和聚集体自噬的NBR1蛋白等也被发现有类似SLRs的作用[11]。如图1-C,除SLRs介导的选择性自噬外,补体C3(complement component C3, C3)被证实为病原体自噬的替代激活物,C3标记细菌后,随细菌进入胞质,直接与ATG16L1(autophagy-related protein 16 like protein 1)结合,刺激宿主病原体自噬并限制胞内细菌增殖[12]。三重基序家族蛋白(family of tripartite motif, TRIM)在自噬和免疫中也具有关键功能,TRIM16在半乳凝素3(galectin 3)的协助下招募ULK1(unc-51-like kinase 1)、BECN1(beclin 1)和ATG16L1操纵转录因子EB(transcriptional factor EB, TFEB)的核转位,激活自噬通路并促进自噬体与溶酶体融合[13]。
-
自噬激活后,被招募的ULK / ATG1激酶复合物、Beclin 1、ATG16L1复合体等自噬相关蛋白(autophagy related protein, ATG)聚集到细菌周围成核,在ATG12类泛素化连接系统(ubiquitin like system, UBL)中,ATG12 作为类泛素蛋白与ATG5共价连接,随后ATG5-ATG12与ATG16L1的N端结合,ATG16L1中部的螺旋卷曲结构域(coiled-coil domain, CCD)二聚化,形成一个约240 ku的ATG5-ATG12-ATG16L1复合物,该复合物发挥类泛素连接酶E3(E3 ubiquitin ligases)的作用,协助磷脂酰乙醇胺(phosphatidylethanolamine, PE)与微管相关蛋白1A / 1B-轻链3(microtubule- associated proteins 1A and 1B, MAP1LC3 / LC3)的羧基末端共价连接,组装、延长吞噬泡的双层膜结构[14]。LC3插入到吞噬体(phagosome)的膜结构中,参与其双层膜的延伸与闭合,促进自噬体的成熟。自噬体成熟后,大部分ATG蛋白脱离双层膜,而被PE修饰的LC3(LC3-PE / LC3-II)仍存在于自噬体的内外膜上,所以,对胞内LC3的检测与示踪是目前常用的检测自噬通路的方法[15]。自噬体成熟后,突触融合蛋白17(syntaxin 17, STX17)定位于其外膜,促进自噬体与溶酶体融合,形成自噬溶酶体降解胞内细菌[16]。
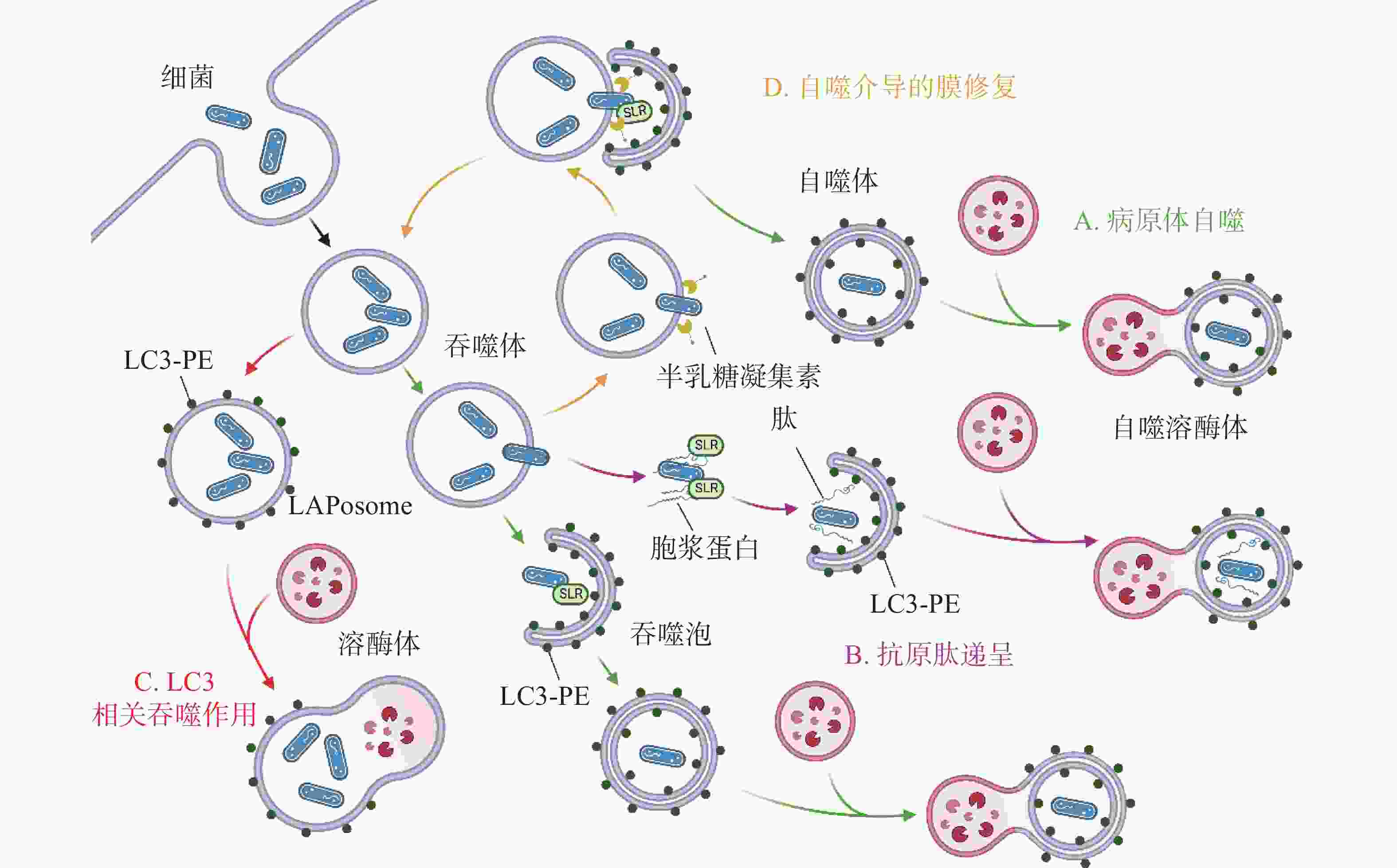
除了典型的病原体(图2-A)激活自噬途径外,胞浆中的抗菌肽(antimicrobial peptides , AMPs),如泛素偶联物(ubiquitin conjugates)和核糖体蛋白(ribosomal proteins),也会协助自噬的激活并将自噬体定位到细菌周围(图2-B),这种现象被成为抗菌肽递呈(antimicrobial peptide delivery)[17]。在中性粒细胞等吞噬细胞中,细菌被TLR或Fc受体配体识别,LC3、Beclin 1、ATG16L1复合体转位到内含细菌的核内体(endosome)膜结构上,促进底物与溶酶体融合(图2-C),这一过程被称为LC3相关吞噬作用(LC3-associated phagocytosis, LAP)。与经典自噬通路不同的是,此时自噬相关蛋白定位的核内体/吞噬体为单层膜结构,其形成需要吞噬细胞NADPH氧化酶产生的活性氧簇(reactive oxygen species, ROS),而不依赖于ULK复合物[18]。值得注意的是,自噬体膜被逃逸的细菌破坏后,暴露在膜表面的多糖被半乳糖凝集素(galectin)识别,一方面能促进病原体自噬,另一方面还会诱发自噬介导的膜修复(membrane repair)功能(图2-D),修复受损的吞噬体膜,延缓细菌逃逸。
-
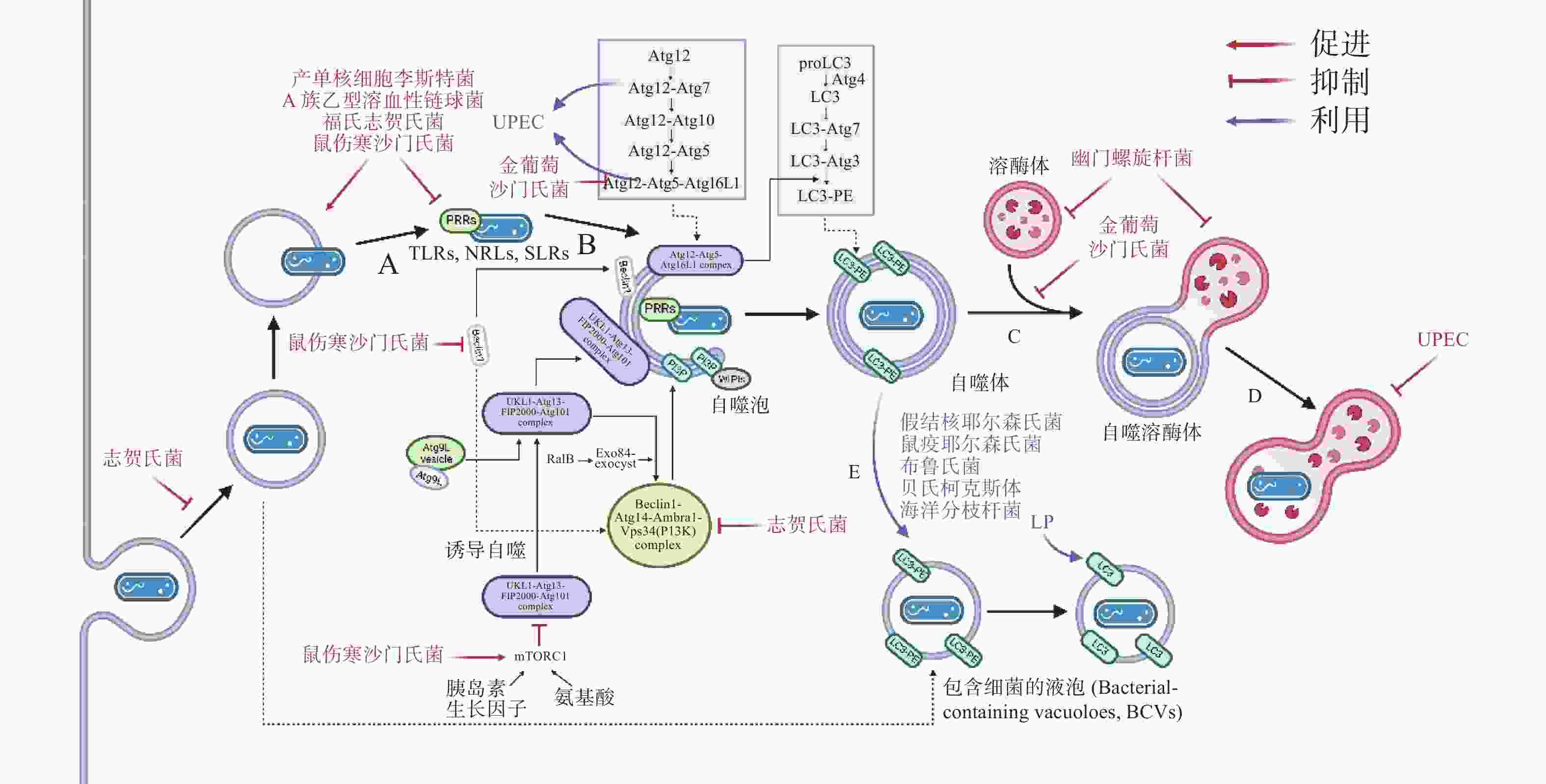
自噬作为新型天然免疫通路,其模式识别受体(pattern recognition receptors, PRRs)在感染后对病原的特异性识别尤为关键。根据遗传进化与系统发育分析,目前细菌公认有Ⅰ-Ⅶ型分泌系统(secretion systems),其分泌的效应蛋白(effectors)可模拟或直接共价修饰宿主蛋白,实现免疫逃逸。
在自噬激活早期,为躲避细胞自噬,细菌分泌效应蛋白抑制PRRs的识别。如图3-A,产单核细胞李斯特菌(Listeria monocytogenes)感染细胞后通过分泌打孔毒素(pore-forming toxin, PFO)、溶血素O(listeriolysin O, LLO)破坏吞噬泡膜,从吞噬泡中逃逸后在细胞质中游离复制[19]。此外,L. monocytogenes表面的ActA蛋白与细胞Arp2/3复合物结合,使微丝蛋白聚集,进而躲避自噬PRRs的识别,当细菌actA基因缺失后,被自噬体包裹的比例明显增加[20]。A族乙型溶血性链球菌(Group A Streptococcus, GAS)分泌的毒力因子SpeB可降解包括SQSTM1 / p62、NDP52和NBR1在内的SLRs,抑制自噬激活[21]。除拮抗SLRs外,鼠伤寒沙门氏菌(Salmonella typhimurium)感染细胞后释放的效应蛋白Sop通过促进TRIM56和TRIM65的泛素化使其降解,从而抑制IFN-γ驱动的自噬[22]。福氏志贺氏菌(Shigella flexneri)分泌的III型效应蛋白IcsB或VirA均可以抑制宿主自噬系统对细菌的识别,且二者独立作用于自噬通路中[23]。类似地,S. typhimurium或S. flexneri分别进化出膜蛋白酶 IcsP或PgtE以摆脱补体C3的标记从而躲避C3驱动的细胞自噬[12]。金黄色葡萄球菌(Staphylococcus aureus)分泌的α溶血素(α-toxin, α-hemolysin)可与ATG16L1及其他一些ATG蛋白介导外泌体(exosomes)释放的ADAM10特异性结合,在降低细胞毒性的同时提高细菌的入胞率[24]。
病原体自噬激活后,其最大特点是形成一个将待降解物质包裹起来的双层膜结构,即自噬体。自噬体的形成是病原体自噬应对胞内细菌感染的重要步骤。为抑制自噬体的形成与成熟,诸多细菌进化出干扰ULK1、ATG5以及ATG16L1等核心ATG的分子机理,如图3-C。S. typhimurium通过III型分泌系统(type III secretion system, T3SS)效应蛋白SrrB促进单磷酸腺苷活化的蛋白激酶(AMP-activated protein kinase, AMPK)/ Sirt1 / LBK1 复合物的降解来破坏AMPK途径的激活,上调mTOR(mechanistic target of rapamycin)水平,抑制ULK复合物的形成[25]。S. aureus进入NIH 3T3 细胞后,刺激MAPK14 / p38对ATG5的磷酸化以抑制自噬体的成熟[26]。Dinic M等[27]发现这一过程也同时抑制了溶酶体与自噬体的融合。沙门氏菌(Salmonella)T3SS 效应器SseF和SseG可破坏GTPase Rab1A的鸟嘌呤核苷酸交换因子(nucleotide exchange factor, GEF)与转运蛋白颗粒III(transport protein particle III, TRAPPIII)的相互作用,这将阻碍ULK1复合物的组装并下调磷脂酰肌醇3-磷酸(phosphatidylinositol 3-phosphate, PI3P)的合成,抑制吞噬泡的成核及膜延伸。最新研究发现,沙门氏菌效应蛋白SopF及其等位效应蛋白通过促进ADP核糖修饰V-ATPase复合物特异抑制ATG16L1的招募,且这一修饰不影响V-ATPase泵H+的能力[28]。
自噬体与溶酶体的融合后,水解酶(hydrolytic enzymes)、酸性磷酸酶(acid phosphatase)、α-葡萄糖苷酶(α-glucosidase)等溶酶体酶分解、消化胞内菌,这些酶在 pH值约为5时活性最佳,因此,抑制自噬体与溶酶体融合、提高自噬溶酶体pH可促进胞内细菌的存活。生理条件下, 6-磷酸甘露糖受体(mannose-6-phosphate receptors, M6PR)负责将合成的水解酶从反式高尔基体网络(trans-Golgi network, TGN)运输到内吞溶酶体[29]。沙门氏菌分泌的效应因子SifA与小鸟苷三磷酸酶 Rab9 结合,阻断M6PR运输的同时降低水解酶的活性[30]。与沙门氏菌类似,幽门螺杆菌(Helicobacter pylori)入侵细胞后,宿主细胞中M6PR逆向运输,溶酶体酸化被抑制[31]。此外,Capurro 等[32]的研究证明,H. pylori产生的空泡毒素A(vacuolating cytotoxin A, VacA)通过靶向溶酶体钙离子通道 TRPML1破坏自噬溶酶体的形成,自噬体即便捕获、包裹了细菌,却无法借助溶酶体酶杀灭它们,同时,这样的胞内小室还为细菌提供了较好的增殖环境(图3-D)。除抑制M6PR的运输,尿路致病性大肠杆菌(uropathogenic Escherichia coli, UPEC)通过提高溶酶体pH值、激活溶酶体特异性钙离子通道逃避自噬降解,该通道使钙离子穿过溶酶体膜到胞质,触发溶酶体的胞吐作用(exocytosis),导致UPEC的排出[33] 。
-
除抑制细胞自噬外,某些胞内寄生菌还能通过诱导、利用自噬促进自身增殖。这些细菌在自噬缺陷细胞中表现出复制缺陷,而用自噬激活剂促进细胞自噬后同时促进了细菌增殖。
自噬为细胞提供基本的营养物质,而在感染情况下,某些细菌入侵细胞后,劫持自噬体或ATG用于自身复制(图3-E)。多数情况下,它们会主动诱导自噬,同时阻止自噬体的成熟以及自噬体与溶酶体的融合。UPEC 通过劫持ATG16L1 和ATG7将铁蛋白上的铁离子释放出来,利用游离的铁离子促进自身的生长[34]。H. pylori 利用胆固醇-葡萄糖苷转移酶(cholesteryl-glucoside transferase)来操纵宿主细胞的胆固醇代谢,将糖基化或脂化的胆固醇用于自身细胞壁合成[35]。此时自噬水平的提高,不仅不能帮助细胞清除细菌,反而促进了细菌对胞内营养物质的获取。此外, Lai等[36]发现,H. pylori对胆固醇的代谢调节抑制了溶酶体与自噬体的融合,敲低ATG12、ATG5以及Beclin1等ATG后抑制了H. pylori的胞内增殖。S. aureus的α毒素(α-toxin)可诱导ATG5依赖性自噬,降低细胞cAMP水平,去除ATG5的细胞中S. aureus载量减少,提示自噬是S. aureus增殖所必需的。
细菌入侵细胞后,在宿主细胞内形成包裹细菌的液泡(bacteria-containing vacuoles),这些液泡为细菌的胞内增殖提供了有利的空间和充足的营养,成为细菌利用细胞自噬的策略之一。嗜肺军团菌(Legionella pneumophila, LP)被细胞吞噬后,在胞内形成包裹军团菌的液泡(Legionella-containing vacuoles, LCVs),从内质网(endoplasmic reticulum, ER)招募囊泡并获得LC3等自噬标志物,使LCVs快速转变为自噬体[37]。这一过程被指依赖于的Ⅳ型分泌系统(Type Ⅳ secretion system, T4SS)效应蛋白LegA9,其通过自噬促进细胞对LCVs的识别,感染2 h后抑制细胞自噬,LP的活率也有所降低,提示LCVs向自噬的转变促进了LP的成活。但是,Choy等[38]随后发现,LP通过分泌LpSPL和RavZ特异性识别LC3-II,并将其C端甘氨酸与酪氨酸之间的酰胺键切割,LC3-II不再能被PE修饰,所有类型的细胞自噬通路被抑制。LP一方面通过LegA9将LCVs靶向自噬,避免被宿主细胞立刻捕获杀死;另一方面借助LpSPL和RavZ抑制自噬,这样的双重策略说明细菌既可以利用细胞自噬实现自身增殖,又通过调节自噬进行免疫逃逸。
除了LP外,其他一些细菌也采用了类似策略,假结核耶尔森氏菌(Yersinia pseudotuberculosis)在胞内形成包裹耶尔森氏菌的液泡(Yersinia-containing vacuoles, YCVs), YCVs上有自噬标志物LC3,雷帕霉素(rapamycin, RAPA)刺激细胞自噬,YCVs大小增大,液泡内细菌载量增加[39]。不仅如此,Pujol等[40]发现,鼠疫耶尔森氏菌(Yersinia pestis)也有在LC3-II修饰的YCVs中复制的特性,他们推测自噬体可能与次级内体(late endosome)一起作为膜来源,促进YCVs的延伸与扩展。布鲁氏菌(Brucella)在ER衍生的包裹布鲁氏菌的液泡(Brucella-containing vacuoles, BCVs)中复制,BCVs劫持ULK1、Beclin1等自噬起始因子,形成类似自噬体的隔室[41]。自噬帮助细菌修复包裹细菌的液泡,在海洋分枝杆菌(Mycobacterium marinum)中更为典型,M. marinum促进自噬相关基因的表达的同时,借助效应蛋白ESX-1阻断自噬通量(autophagic flux), 通过促进自噬修复包裹分支杆菌的液泡(Mycobacteria-containing vacuoles, MCVs)[42]。
-
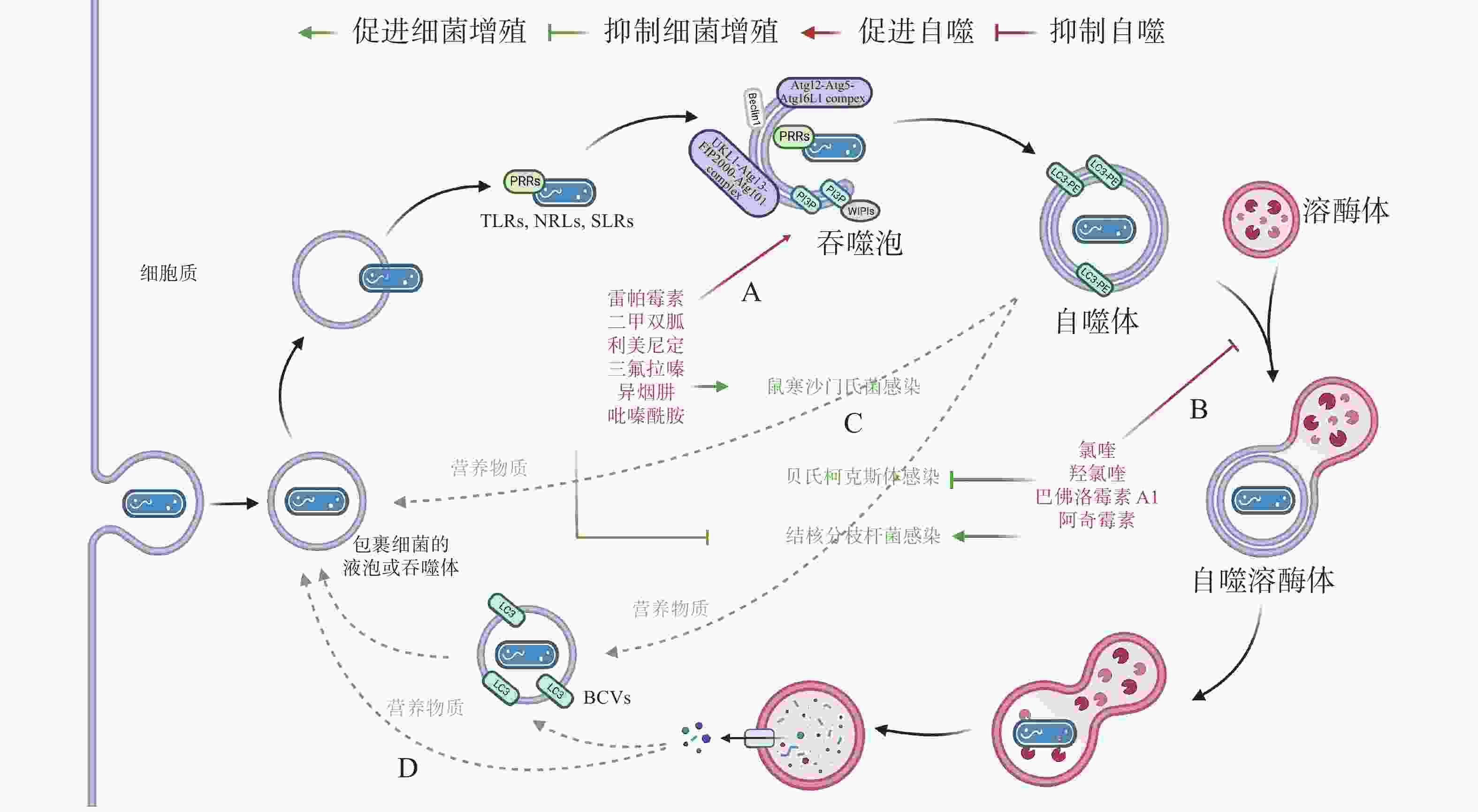
作为新型天然免疫通路,借助细胞自噬控制胞内细菌的增殖可成为抗生素治疗的替代途径,自噬调节剂不仅能于不同阶段调节自噬,还能通过促进或抑制自噬治疗感染性疾病。目前,已有一些自噬调节剂于临床上广泛使用(图4-A):RAPA、二甲双胍(metformin)和利美尼定(rilmenidine)等自噬诱导剂用于预防肾移植排斥或治疗2型糖尿病和高血压等[43];氯喹(chloroquine, CQ)和羟氯喹(hydroxychloroquine, HCQ)等抗疟疾药物目前正作为自噬拮抗剂在临床试验中用于治疗某些耐药癌症[44]。此外,HCQ联合强力霉素(doxycycline)被用于治疗C. burnetii引起的慢性Q热(Q fever)心内膜炎[45]。
有研究发现,异烟肼(isoniazid, INH)和吡嗪酰胺(pyrazinamide, PZA)等是广泛用于治疗结核分枝杆菌(Mycobacterium tuberculosis)的药物,它们不仅能直接杀死细菌,还能通过诱导细胞自噬,促进宿主对胞内M. tuberculosis的清除[46]。阿奇霉素(azithromycin, AZI)通过抑制溶酶体酸化阻断M. tuberculosis自噬降解,长期使用AZI可能导致条件性分枝杆菌感染[47]。INH 或 PZA 等自噬诱导剂可成功治疗结核病,而AZI 等自噬抑制剂促进M. tuberculosis感染,表明自噬是细胞对抗M. tuberculosis感染的有效途径之一。与M. tuberculosis类似,Conway等[48]发现三氟拉嗪(trifluoperazine)可抑制S. typhimurium在HeLa 细胞中的复制,而这种抑制作用是ATG16L1依赖的,表明利用细胞自噬可促进胞内细菌的清除(图4-C)。
药物调节自噬可作为对抗胞内细菌感染的候选策略,促进细胞自噬在应对一些细菌入侵时效果显著,但如(图4-D)所示,UPEC、H. pylori和LP等细菌利用细胞自噬促进自身生长,所以,在使用自噬调节剂治疗传染性疾病时不仅需要高度谨慎,在治疗过程中还须实时监测感染风险。
-
细菌感染诱发的细胞自噬作为天然免疫及细胞代谢的一部分,是细胞内细菌清除与增殖之间的博弈,有着复杂的分子机理和重要的生物学功能。在细菌入侵过程中,一方面细胞通过自噬途径清除胞内病原体,另一方面细菌借助自噬获得胞内营养。目前,细胞如何借助自噬系统识别、清除细菌,细菌进化出了哪些自噬逃逸机制尚不完全明了。今后,随着更多的自噬相关蛋白及影响自噬通路的细菌蛋白的发现,人们对自噬体识别细菌机制的认识会更加深入。同时,自噬与细胞其他信号通路之间存在着复杂的关联,简单地聚焦于细胞自噬与细菌感染的相互作用不能完全解释胞内细菌载量的变化,需要进一步探索各天然免疫通路之间的窜扰(crosstalk),开展相关研究。
通过研究认识自噬在天然免疫系统中的作用,明确细菌逃逸、利用细胞自噬的分子机理,对于利用自噬调节对抗细菌感染、开发针对细胞内细菌感染的新药靶点、构建新的细菌防控策略尤为重要。但值得注意的是,细菌感染诱发细胞自噬具有两面性,在用药治疗的过程中,须高度谨慎,并在治疗过程中实时监测。
Advances in autophagy induced by bacterial infection
-
摘要: 细胞内的细菌清除和增殖是细菌感染和细胞自噬之间的博弈:细胞借助自噬系统识别、清除胞内细菌;细菌进化出不同的逃逸机制,抑制、利用细胞自噬,促进自身增殖。针对细菌感染与细胞自噬之间的复杂关系,对细菌激活细胞自噬途径、自噬反应通路、细菌与自噬的相互作用以及感染性疾病治疗中的自噬调节进行了总结,综述了现阶段病原体自噬及细菌逃逸、利用细胞自噬的研究进展。Abstract: The autophagy pathway activated by bacteria has two faces in maintaining the dynamic balance between intracellular bacterial removal and proliferation. On the one hand, cells recognize and remove intracellular bacteria via autophagy; on the other hand, bacteria evolve various escape mechanisms to inhibit and utilize autophagy to promote their own proliferation. A comprehensive summary was made of the complex relationship between bacterial infection and autophagy, involving the activation of autophagic pathways by bacteria, autophagic response pathways, the interaction between bacteria and autophagy, and the regulation of autophagy in the treatment of infectious diseases to review recent advances in bacterial infection and autophagy, as well as the utilization of autophagy, so as to provide reference for subsequent autophagy exploration.
-
Key words:
- autophagy /
- bacterial infection /
- immune escape
-
-
[1] KUMA A, KOMATSU M, MIZUSHIMA N. Autophagy-monitoring and autophagy-deficient mice [J]. Autophagy, 2017, 13(10): 1619 − 1628. doi: 10.1080/15548627.2017.1343770 [2] 张宏, 张慧. 多细胞生物自噬的分子机制和生理功能[J]. 安徽大学学报(自然科学版), 2018, 42(5): 105 − 114. [3] KWON D H, SONG H K. A Structural view of xenophagy, a battle between host and microbes [J]. Mol Cells, 2018, 41(1): 27 − 34. [4] FENG Y, HE D, YAO Z, et al. The machinery of macroautophagy [J]. Cell Res, 2014, 24(1): 24 − 41. doi: 10.1038/cr.2013.168 [5] LI W W, LI J, BAO J K. Microautophagy: lesser-known self-eating [J]. Cell Mol Life Sci, 2012, 69(7): 1125 − 1136. doi: 10.1007/s00018-011-0865-5 [6] DICE J F. Chaperone-mediated autophagy [J]. Autophagy, 2007, 3(4): 295 − 299. doi: 10.4161/auto.4144 [7] WANG Z, LI C. Xenophagy in innate immunity: A battle between host and pathogen [J]. Dev Comp Immunol, 2020, 109: 103693. doi: 10.1016/j.dci.2020.103693 [8] LI S, HE J, XU H, et al. Autophagic activation of IRF-1 aggravates hepatic ischemia-reperfusion injury via JNK signaling [J]. MedComm, 2021, 2(1): 91 − 100. [9] DERETIC V. Autophagy as an innate immunity paradigm: expanding the scope and repertoire of pattern recognition receptors [J]. Curr Opin Immunol, 2012, 24(1): 21 − 31. doi: 10.1016/j.coi.2011.10.006 [10] LEE Y J, KIM J K, JUNG C H, et al. Chemical modulation of SQSTM1/p62-mediated xenophagy that targets a broad range of pathogenic bacteria [J]. Autophagy, 2022, 18(12): 2926 − 2945. doi: 10.1080/15548627.2022.2054240 [11] TURCO E, SAVOVA A, GERE F, et al. Reconstitution defines the roles of p62, NBR1 and TAX1BP1 in ubiquitin condensate formation and autophagy initiation [J]. Nat Commun, 2021, 12(1): 5212. doi: 10.1038/s41467-021-25572-w [12] MATTHEW T S, ELISABETH G F, JESSICA T, et al. Complement C3 drives autophagy-dependent restriction of cyto-invasive bacteria [J]. Cell Host & Microbe, 2018, 23(5): 644 − 652. [13] DI RIENZO M, ROMAGNOLI A, ANTONIOLI M, et al. TRIM proteins in autophagy: selective sensors in cell damage and innate immune responses [J]. Cell Death Differ, 2020, 27(3): 887 − 902. doi: 10.1038/s41418-020-0495-2 [14] FISCHER T D, WANG C, PADMAN B S, et al. STING induces LC3B lipidation onto single-membrane vesicles via the V-ATPase and ATG16L1-WD40 domain [J]. J Cell Biol, 2020, 219(12): .1 − 17. [15] INOMATA M, XU S, CHANDRA P, et al. Macrophage LC3-associated phagocytosis is an immune defense against Streptococcus pneumoniae that diminishes with host aging [J]. Proc Natl Acad Sci U S A, 2020, 117(52): 33561 − 33569. doi: 10.1073/pnas.2015368117 [16] NOWACKA-WOSZUK J, MACKOWSKI M, MANTAJ W, et al. Equine STX17 intronic triplication confirmed by droplet digital PCR analysis of its breakpoints [J]. Anim Genet, 2021, 52(4): 567 − 568. doi: 10.1111/age.13073 [17] GAO N, YANG Y, LIU S, et al. Gut-derived metabolites from dietary tryptophan supplementation quench intestinal inflammation through the AMPK-SIRT1-autophagy pathway [J]. J Agric Food Chem, 2022, 70(51): 16080 − 16095. doi: 10.1021/acs.jafc.2c05381 [18] PRAJSNAR T K, SERBA J J, DEKKER B M, et al. The autophagic response to Staphylococcus aureus provides an intracellular niche in neutrophils [J]. Autophagy, 2021, 17(4): 888 − 902. doi: 10.1080/15548627.2020.1739443 [19] TINGTING W, TIANQI F, XINYU W, et al. Amentoflavone attenuates Listeria monocytogenes pathogenicity through an LLO-dependent mechanism [J]. Br J Pharmacol, 2022, 179(14): 3839 − 3858. doi: 10.1111/bph.15827 [20] BOUJEMAA-PATERSKI R, GOUIN E, HANSEN G, et al. Listeria protein ActA mimics WASp family proteins: it activates filament barbed end branching by Arp2/3 complex [J]. Biochemistry, 2001, 40(38): 11390 − 11404. doi: 10.1021/bi010486b [21] LIN C Y, NOZAWA T, MINOWA N A, et al. Autophagy receptor tollip facilitates bacterial autophagy by recruiting galectin-7 in response to Group A streptococcus infection[J]. Frontiers in Cellular and Infection Microbiology, 2020(10). doi: 10.3389/fcimb.2020.583137. [22] BIRMINGHAM C L, BRUMELL J H. Autophagy recognizes intracellular Salmonella enterica serovar Typhimurium in damaged vacuoles [J]. Autophagy, 2006, 2(3): 156 − 158. doi: 10.4161/auto.2825 [23] CAMPBELL-VALOIS F X, SACHSE M, SANSONETTI P J, et al. Escape of actively secreting Shigella flexneri from ATG8/LC3-Positive vacuoles formed during cell-to-cell spread is facilitated by IcsB and VirA [J]. MBio, 2015, 6(3): 2514 − 2567. [24] KELLER M D, CHING K L, LIANG F X, et al. Decoy exosomes provide protection against bacterial toxins [J]. Nature, 2020, 579(7798): 260 − 264. doi: 10.1038/s41586-020-2066-6 [25] MA L, LI W, ZHANG Y, et al. FLT4/VEGFR3 activates AMPK to coordinate glycometabolic reprogramming with autophagy and inflammasome activation for bacterial elimination [J]. Autophagy, 2022, 18(6): 1385 − 1400. doi: 10.1080/15548627.2021.1985338 [26] JIN T, HE P, YANG R, et al. CHI3L1 promotes Staphylococcus aureus-induced osteomyelitis by activating p38/MAPK and Smad signaling pathways [J]. Exp Cell Res, 2021, 403(1): 112596. doi: 10.1016/j.yexcr.2021.112596 [27] DINIC M, JAKOVLJEVIC S, DOKIC J, et al. Probiotic-mediated p38 MAPK immune signaling prolongs the survival of Caenorhabditis elegans exposed to pathogenic bacteria [J]. Sci Rep, 2021, 11(1): 21258. doi: 10.1038/s41598-021-00698-5 [28] XU Y, CHENG S, ZENG H, et al. ARF GTPases activate Salmonella effector SopF to ADP-ribosylate host V-ATPase and inhibit endomembrane damage-induced autophagy [J]. Nat Struct Mol Biol, 2022, 29(1): 67 − 77. doi: 10.1038/s41594-021-00710-6 [29] CHANG F, LI N, YAN K, et al. Luminal/extracellular domains of chimeric CI-M6PR-C proteins interfere with their retrograde endosome-to-TGN trafficking in the transient expression system [J]. J Biomed Res, 2018, 32(4): 245 − 256. doi: 10.7555/JBR.32.20180044 [30] RABINOVICH-NIKITIN I, COGAN R C, KIRSHENBAUM L A. Attenuation of obesity cardiomyopathy by Ulk1/Rab9 mediated alternative mitophagy [J]. Circ Res, 2021, 129(12): 1122 − 1124. doi: 10.1161/CIRCRESAHA.121.320365 [31] JALAGADUGULA G, MAO G, GOLDFINGER L E, et al. Defective RAB31-mediated megakaryocytic early endosomal trafficking of VWF, EGFR, and M6PR in RUNX1 deficiency [J]. Blood Adv, 2022, 13,(17): 5100 − 5112. [32] CAPURRO M I, PRASHAR A, JONES N L. MCOLN1/TRPML1 inhibition - a novel strategy used by Helicobacter pylori to escape autophagic killing and antibiotic eradication therapy in vivo [J]. Autophagy, 2020, 16(1): 169 − 170. doi: 10.1080/15548627.2019.1677322 [33] VASUDEVAN S, THAMIL S G, BHASKARAN S, et al. Reciprocal cooperation of Type A procyanidin and nitrofurantoin against multi-drug resistant (MDR) UPEC: A pH-dependent study [J]. Front Cell Infect Microbiol, 2020, 10: 421. doi: 10.3389/fcimb.2020.00421 [34] 段灵涛, 祝一鸣, 何九卿, 等. 真菌细胞自噬的研究进展[J]. 热带生物学报, 2021, 12(2): 253 − 260. doi: 10.15886/j.cnki.rdswxb.2021.02.015 [35] JAN H M, CHEN Y C, YANG T C, et al. Cholesteryl alpha-D-glucoside 6-acyltransferase enhances the adhesion of Helicobacter pylori to gastric epithelium [J]. Commun Biol, 2020, 3(1): 120. doi: 10.1038/s42003-020-0855-y [36] LAI C H, HUANG J C, CHENG H H, et al. Helicobacter pylori cholesterol glucosylation modulates autophagy for increasing intracellular survival in macrophages [J]. Cell Microbiol, 2018, 20(12): e12947. doi: 10.1111/cmi.12947 [37] OMOTADE T O, ROY C R. Legionella pneumophila excludes autophagy adaptors from the ubiquitin-labeled vacuole in which it resides [J]. Infect Immun, 2020, 88(8): e00793 − 19. doi: 10.1128/IAI.00793-19 [38] CHOY A, ROY C R. Autophagy and bacterial infection: an evolving arms race [J]. Trends Microbiol, 2013, 21(9): 451 − 456. doi: 10.1016/j.tim.2013.06.009 [39] LEMARIGNIER M, PIZARRO-CERDA J. Autophagy and intracellular membrane trafficking subversion by pathogenic Yersinia species [J]. Biomolecules, 2020, 10(12): 1637. doi: 10.3390/biom10121637 [40] PUJOL C, KLEIN K A, ROMANOV G A, et al. Yersinia pestis can reside in autophagosomes and avoid xenophagy in murine macrophages by preventing vacuole acidification [J]. Infect Immun, 2009, 77(6): 2251 − 2261. doi: 10.1128/IAI.00068-09 [41] ZHANG L, YU S, NING X, et al. A LysR transcriptional regulator manipulates macrophage autophagy flux during brucella infection [J]. Front Cell Infect Microbiol, 2022, 12: 858173. doi: 10.3389/fcimb.2022.858173 [42] MUNOZ-SANCHEZ S, VAN DER VAART M, MEIJER A H. Autophagy and Lc3-associated phagocytosis in zebrafish models of bacterial infections [J]. Cells, 2020, 9(11): 2372. doi: 10.3390/cells9112372 [43] GUNAWAN M, LOW C, NEO K, et al. The role of autophagy in chemical proteasome inhibition model of retinal degeneration [J]. Int J Mol Sci, 2021, 22(14): 7271. doi: 10.3390/ijms22147271 [44] DEVIS-JAUREGUI L, ERITJA N, DAVIS M L, et al. Autophagy in the physiological endometrium and cancer [J]. Autophagy, 2021, 17(5): 1077 − 1095. doi: 10.1080/15548627.2020.1752548 [45] VAN ROEDEN S E, BLEEKER-ROVERS C P, DE REGT M, et al. Treatment of chronic Q fever: Clinical efficacy and toxicity of antibiotic regimens [J]. Clin Infect Dis, 2018, 66(5): 719 − 726. doi: 10.1093/cid/cix886 [46] GUERRERO-BUSTAMANTE C A, DEDRICK R M, GARLENA R A, et al. Toward a phage cocktail for tuberculosis: Susceptibility and tuberculocidal action of mycobacteriophages against diverse mycobacterium tuberculosis strains [J]. mBio, 2021, 12(3): 921 − 937. doi: 10.1128/mBio.00973-21 [47] RENNA M, RUBINSZTEIN D C. Macroautophagy without LC3 conjugation? [J]. Cell Res, 2017, 27(1): 5 − 6. doi: 10.1038/cr.2016.143 [48] CONWAY K L, KUBALLA P, SONG J H, et al. Atg16l1 is required for autophagy in intestinal epithelial cells and protection of mice from Salmonella infection [J]. Gastroenterology, 2013, 145(6): 1347 − 1357. doi: 10.1053/j.gastro.2013.08.035 -



 下载:
下载: