-
盐碱地是中国主要的退化土壤类型之一,其改良与治理是开发后备土地资源、提升土地质量的重要途径[1-2]。中国盐碱地总面积近 1 亿 hm2,受盐碱化危害的耕地 920.94×104 hm2 [3]。同时,重金属污染土壤是中国耕地土壤污染的重要类型,耕地土壤重金属超标率为 35% 左右[4]。盐碱化与重金属污染在空间上常重叠发生,形成更为复杂、顽固的复合污染,对传统修复技术构成严峻挑战。目前二者复合污染虽无统一的官方统计数据,但其实际分布较广,亟待探索能同步适应高盐与重金属双重胁迫的绿色修复方案。
重金属胁迫是危害性极强的非生物胁迫。铅、镉、汞、砷等重金属因其高毒性和不可降解性[5],已通过食物链威胁生态安全与人类健康。据最新研究估计,全球约14%~17%的耕地土壤至少一种重金属含量超标,受影响面积达2.42亿hm2;其中镉污染最为普遍,全球耕地超标率约为9.0%[6]。虽然植物需要微量的重金属如钴、铜、铁、锰、钼、镍、钒和锌参与代谢,但过量重金属会竞争必需养分,干扰离子稳态并诱导氧化损伤,严重抑制植物生长[7]。重金属污染的修复引起了研究和生产实际的重视,传统的物理化学修复技术因成本高、劳动强度大、易破坏土壤结构而受限,已难以满足大面积治理的需求。相比之下,植物修复技术以低成本、环境友好等突出优势,成为重金属污染治理的重要可行途径。
然而,普通超富集植物(hyperaccumulator plants)在盐碱环境中的适应性有限,表现为生物量低、耐盐性差等缺陷,极大地制约了其在盐碱-重金属复合污染场地的应用。盐生植物(halophytes)则因其原生耐盐性,展现出独特的优势。凭借其独特的重金属-盐度交叉耐受性(crossed tolerance),能够在高盐条件下仍保持对重金属的高效吸收与转运。其根系可分泌有机酸、调控重金属氧化还原态(如Fe3+/Fe2+转化),并与根际微生物协同作用,显著降低重金属生物有效性。此外,与普通植物相比,盐生植物拥有高效的离子转运系统,如重金属ATP酶(heavy metal ATPase,HMA)、天然抗性相关巨噬细胞蛋白(natural resistance-associated macrophage protein,NRAMP)家族蛋白,这些蛋白能够将根部吸收的金属快速转运至地上部,降低根部细胞毒性并提升整体耐受水平[8]。这种交叉耐受性不仅是一种生理现象,其背后蕴含着共享的分子调控基础,例如,共用的信号通路[如促分裂原活化蛋白激酶(mitogen-activated protein kinase,MAPK)级联]、协同的激素信号网络[如脱落酸(abscisic acid, ABA)和茉莉酸(jasmonic acid, JA)],以及表观遗传调控机制(如DNA甲基化)。目前,研究者已通过分子生物学手段解析植物中金属吸收、转化及耐受机制[9],但盐生植物交叉耐受性的系统性分子机制仍有待深入阐明。鉴于盐生植物能够通过多种转运蛋白将摄取的金属离子从根部输送至地上部,深入探讨其在重金属等非生物胁迫下的分子水平耐受机制,不仅有助于阐明盐生植物适应极端环境的生存策略,更能为挖掘其植物修复潜力,进而通过分子设计育种定向选育兼具高耐盐性、强重金属富集能力的“设计型”盐生修复植物,提供关键的理论靶标和基因资源。本文将系统综述盐生植物对重金属的耐受性及其分子机制,重点剖析其与耐盐机制共享的分子基础(交叉耐受性),并着重探讨如何将这些分子层面的认知(如关键基因、信号通路、微生物互作)与分子标记辅助选择、基因工程育种等现代选育策略相结合。最终,旨在从“分子机制-选育应用”衔接的视角,为盐碱地-重金属复合污染的绿色、可持续治理,以及盐生植物种质资源的创新利用,提供兼具理论依据和应用导向的参考。
-
盐生植物在长期进化中形成了多种重金属的适应策略,按作用机制可分为三类典型模式:假盐生植物(recretohalophytes)根际离子的排斥作用,通过限制金属进入木质部,显示出排斥或避免机制[10],如海榄雌(Avicennia marina);泌盐盐生植物(excretohalophytes)借助盐腺或盐囊泡主动排出过量离子,木榄(Avicennia marina)能有效地从根部排除铜离子和镉离子,并且对这些重金属表现出高耐受性[11];真盐生植物(euhalophytes),如盐地碱蓬(Suaeda salsa)和盐角草(Salicornia europaea),虽然缺乏泌盐结构或根系离子排斥能力,但其能通过分子适应性机制应对盐与重金属复合胁迫,为防止盐毒离子在细胞质中积累,它们会被隔离在液泡中,实现离子积累[12]。上述盐生植物的分类模式反映了盐生植物应对盐胁迫的策略多样性。同时,与盐离子积累类似,海蓬子属(Salicornia plants)、互花米草(Spartina alterniflora)、海马齿(Sesuvium portulacastrum)均能在地上部分富集重金属离子[13-14]。对于积累过量重金属离子所引起的系列的毒害反应,盐生植物的适应机制涉及不同层面,在植物分子水平主要包括以下方面:
-
活性氧(ROS)产生是生物应对胁迫的共性响应,主要源于叶绿体、线粒体以及过氧化物酶体。重金属积累会损伤叶绿体光反应相关组件功能,降低光合作用电子传递链效率,进而引发ROS积累[15]。线粒体将总耗氧量的1%~5%转化为ROS[16]。在线粒体的光呼吸过程中,乙醇酸(C2H4O3)氧化成乙醛酸(C2H2O3)产生过氧化氢(H2O2)。因此,不同的活性氧分子如单线态氧、超氧阴离子(O2-)、过氧化氢和羟基离子在效应因子触发的易感性(effector-triggered susceptibility,ETS)作用过程中形成[17]。铜、镉、铁、锌等金属通过哈伯-韦斯反应(Haber-weiss)和芬顿反应(Fenton reaction)阻碍抗氧化防御酶的活性,诱导ROS产生[18],ROS在植物代谢中具有双重作用。适宜浓度下可作为胁迫传感器,参与细胞生长、分化,根须生长及气孔形成等生理过程[19]。而过量的ROS会直接与细胞组分(脂质、蛋白质、核酸)发生反应,导致脂质过氧化,膜损伤以及酶失活,影响细胞的正常生命活动[20]。ROS 的稳态受清除系统机制的调控。如果清洁系统能有效清除ROS,低浓度ROS作为信号分子调控基因表达(如ROS信号通路激活胁迫响应基因)。过量ROS导致脂质过氧化、酶失活和细胞损伤,那么ROS就会变成有毒物质[21]。
植物抗氧化系统是清除ROS的核心,由酶类与非酶类成分组成,二者协同保护细胞免受ROS损伤。酶促抗氧化剂是由超氧化物歧化酶(superoxide dismutase,SOD)、过氧化氢酶(catalase,CAT)、过氧化物酶(peroxidase,POD)、抗坏血酸过氧化物酶(aseorbate peroxidase,APX)、谷胱甘肽还原酶(gluathione reductase,GR)等组成。这些酶通过催化反应将活性氧转化为无害产物(如水或无机离子),从而缓解金属胁迫下的氧化损伤。例如,SOD通过歧化反应将O2−转化为H2O2,CAT再将H2O2分解为H2O和O2。此外,非酶类成分如抗坏血酸(ASA)和谷胱甘肽(GSH)通过直接清除ROS或作为抗氧化酶的辅因子(如APX和GR)发挥作用[22]。在重金属胁迫下,植物通过上调抗氧化酶活性以应对ROS的积累。例如,铜处理可显著提高苔藓植物中的过氧化物酶(POD)和抗坏血酸过氧化物酶(APX)活性,而铅胁迫则显著增加SOD、POD、愈创木酚过氧化物酶(guaiacol Peroxidase,GPX)和APX的活性,且其活性随金属浓度升高而增强。这种酶活性的上调可能通过ROS信号通路(如ROS信号通路激活胁迫响应基因)调控,从而增强抗氧化防御[23]。在重金属胁迫缓解中,抗氧化系统与金属适应机制协同作用。例如,GSH不仅作为抗氧化剂清除ROS,还可作为植物螯合肽(PCs)的前体,通过螯合重金属离子(如Cd、Pb)减少其生物毒性[24]。此外,硅(Si)胁迫可增强植物抗氧化系统的活性,通过刺激SOD、CAT等酶的活性缓解重金属胁迫。
植物抗氧化系统是应对重金属胁迫的第一道防线,其关键酶(SOD、CAT、APX)的活性与基因表达水平可作为盐生植物重金属耐受性的重要生理指标。该系统通过酶促和非酶促机制协同清除ROS,维持细胞稳态,其活性变化与金属胁迫的强度和类型密切相关[25]。利用转录组分析或基因编辑技术调控抗氧化基因表达,是提升植物在复合污染环境中生存与修复能力的潜在育种策略。
-
螯合作用(chelation)是盐生植物抵御重金属胁迫的关键机制。植物通过合成特定螯合物质,与体内过量重金属结合形成复合物,实现重金属的运输、区域化储存及胞质离子稳态维持,这也是其缓解离子胁迫损伤的核心途径[26]。在盐生植物的螯合体系中,巯基化合物是研究最为深入且作用关键的一类,这类物质主要包含两大关键成员——植物螯合肽(phytochelatins,PCs)与金属硫蛋白(metallothionein,MT)。二者同属巯基化合物,但其在合成机制、功能特征及调控模式等方面存在差异,具体如下:
-
植物螯合肽(phytochelatins,PCs)是植物螯合酶以GSH为底物合成的低分子量多肽,对重金属具有高亲和力,其可与重金属结合形成稳定复合物以降低毒性,再经转运蛋白转运至液泡中区室化储存,最终维持胞质重金属稳态与植物正常生理功能。在刚毛藻(Cladophora)中,随着镉胁迫程度上升,PCs中镉含量逐渐增加,成为主要的抗胁迫因子[27]。然而,非重金属富集植物中,组成型PCs的基础表达也可能帮助其缓解高金属载荷造成的损伤[28]。在黑海榄雌(Avicennia germinans)中,镉离子与铜离子胁迫可诱导AvPCS基因在6 h内显著上调[29]。尽管该基因表达呈现瞬时激活特征,但其通过级联信号传导可触发持久的重金属适应机制。研究显示,PCs在植物对重金属胁迫的耐受性中起关键作用。例如海滨藜(Atriplex maximowicziana)和碱蓬属(Suaeda spp.)叶片组织中,PCs的过量合成与其对镉、锌胁迫的高耐受性显著相关[30]。
-
与PCs不同,金属硫蛋白(metallothionein,MT)作为另一类关键巯基化合物,结构与功能具有更强的特异性,是一类广泛存在于自然界且富含半胱氨酸残基(Cys)的金属结合蛋白[31]。其结构特性包括半胱氨酸残基的排列方式,可以分为Type 1、Type 2、Type 3、Type 4四种类型[32]。而这四种类型的MT在植物体中往往存在组织特异性表达。在拟南芥中,MT1主要在根部表达,MT2和MT3主要在叶中表达,MT4仅在种子中表达。植物MT基因的表达往往受过量重金属胁迫诱导。例如,在镉胁迫下,浮萍(Spirodela polyrhiza)的SpMT2a基因的转录丰度显著上调,且其异源表达赋予了酵母对铜和镉胁迫的耐受性[33]。这证实了MT基因在重金属响应中的诱导表达特征及功能保守性。在金属胁迫下,植物MT的表达和功能表现出显著的物种和金属特异性。例如,在木榄中,MT2基因在镉、铜和锌胁迫下表现出高表达,表明其在重金属耐受中的关键作用;类似地,在海榄雌中,AmMT-2基因也表现出对镉、铜、铅和锌的高耐受性[34];在牧豆树(Prosopis juliflora)中,PJMT-2基因在锌处理下上调数倍,而在镉和铜处理下其水平基本保持不变[35]。在相同的胁迫处理下,海蓬子(Salvinia biloba)的SbMT-2基因在锌和铜的处理下上调,但是在镉处理下却不会发生变化[36]。表明不同盐生植物的MT基因对重金属离子具有显著的识别与响应特异性。这种特异性本质上由MT基因的启动子调控元件(如金属应答元件、激素应答元件)和转录因子(如MTF-1)的结合密切相关。MT在植物修复和抗逆性中的应用潜力显著。例如,过表达MT基因可提高植物对重金属(如镉、锌)的耐受性,并增强抗氧化酶活性(如SOD、CAT)[37]。此外MT的金属螯合能力(通过半胱氨酸残基)和抗氧化功能(清除活性氧)使其在重金属耐受和非生物胁迫响应中发挥重要作用。
因此,PCs和MT基因是基因工程改良的优选靶点。过表达这些基因,能够强化植物对重金属的螯合与区室化能力,显著提升其富集与耐受水平。针对不同重金属污染土壤,筛选或改造具有特异性金属结合能力的MT/PCs基因变体,将有助于实现定制化的植物修复。
-
盐生植物通过根部吸收金属离子,经特定转运蛋白介导完成根-芽长距离运输,同时将过量重金属转运至液泡区室化储存,进而维持胞质离子稳态。参与这一过程的蛋白主要包括重金属ATP酶(HMA)、天然抗性相关巨噬细胞蛋白(NRAMP)、阳离子扩散促进因子(Cation diffusion facilitator family,CDF)和锌铁转运蛋白(Zinc-regulated transporter, Iron-regulated transporter-like Protein,ZIP)等。
HMA是介导重金属跨膜转运的核心蛋白,可介导Cd2+、Cu2+等二价阳离子的根-芽运输及液泡储存。与非超富集植物相比,HMA家族基因在超富集植物的根和芽中高度上调[32]。除典型转运蛋白家族外,部分特有蛋白也参与金属胁迫响应,如四翅滨藜(Atriplex canescens)ACHMA1蛋白的表达显著提高了酵母对于铁及其他金属的胁迫耐受性[38]。盐生植物冰叶日中花(Mesembryanthemum crystallinum)中克隆得到的阳离子转运蛋白基因McHKT1,其编码的蛋白序列与已知植物(如小麦、水稻等)HKT1类钾(K+)/钠(Na+)转运蛋白具有41–61%的相似性;在非洲爪蟾(Xenopus laevis)卵母细胞中,该蛋白对铷(Rb+)的结合特异性显著高于铯(Cs+),且对Rb+和Cs+的选择性显著高于K+、Na+及锂(Li+)[39-40]。天然抗性相关巨噬细胞蛋白(NRAMP)由多基因家族编码,广泛参与植物根部重金属的吸收与分配。例如水稻(Oryza sativa)中已鉴定出三个Nramp家族基因OsNramp1、OsNramp2和OsNramp3;芥属植物(Brassica juncea)鉴定出五个成员BjNramp1至BjNramp5。这些基因编码的NRAMP蛋白(如OsNramp1、BjNramp1等)因其在金属转运中的功能,常在根部和地上部高度表达[41]。阳离子扩散促进因子(CDF)家族作为一类负责转运重金属离子的新型蛋白质家族,其成员金属耐受蛋白(metal tolerance protein,MTP)参与多种二价阳离子的区室化隔离、螯合与跨膜转运。例如镉胁迫下,千穗谷(Amaranthus hypochondriacus L.)多个AhMTP家族基因呈现组织特异性上调表达,与植株镉耐受性密切相关[42]。ZIP家族蛋白则以转运Zn2+、Fe2+为主,其中锌转运蛋白ZNT1(Zinc Transporter 1)在根与芽组织的高表达可促进Zn2+转运[43],表明植物可通过根-茎转运系统实现金属离子长距离运输,同时维持根部正常生理功能。
盐生植物对重金属的适应,依赖于活性氧清除、螯合解毒与转运区室化三重分子机制构成的多层次协同网络。这一集成化的分子适应性,正是盐生植物应对单一重金属胁迫的核心,也为其适应更为复杂的多重胁迫环境奠定了内在基础。
-
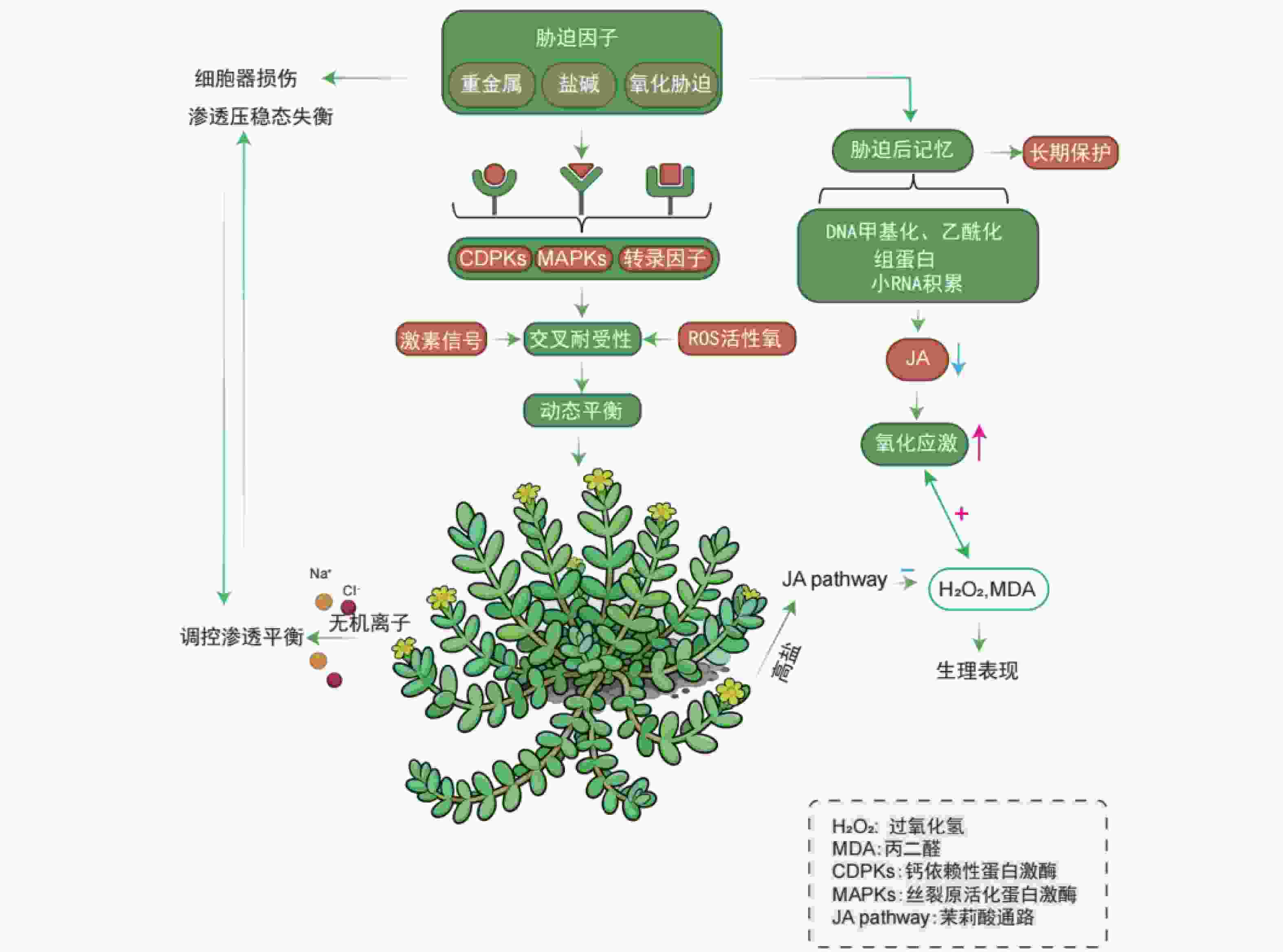
植物经某一生物或非生物胁迫诱导后激活的生理与分子响应,不仅能增强植物自身对该胁迫的适应性,还可提升对后续其他类型胁迫的抗性,这一生物学现象被称为交叉耐受性(cross-tolerance),是植物适应多变环境的关键生存策略[44-45]。盐生植物在长期应对非生物胁迫(如盐害、重金属污染)过程中进化出了交叉耐受性,形成了对生物胁迫的适应能力[46]。交叉耐受性中,不同的信号通路可被同一上游信号事件激活,既可独立引发相似的生理反应,也可通过信号交互整合,共同调控最终的应激响应[47]。这些信号通路可能作为调控因子,在胁迫响应中发挥抑制或促进的双重作用[48-49]。盐生植物在极端生境中可协同耐受高盐与氧化胁迫,其多重胁迫响应机制的核心是信号通路的交叉对话(crosstalk)。该调控网络涉及多胁迫共用的核心信号元件,包括:胁迫感受器、钙依赖性蛋白激酶(Calcium-dependent protein kinase,CDPKs)、丝裂原活化蛋白激酶(Mitogen-activated protein kinase,MAPKs)级联系统及胁迫响应转录因子[50]。进化形成的整合信号系统通过协调激素信号、ROS与抗氧化剂动态平衡,实现有限抗逆资源的最优配置[51]。
重金属胁迫可引发细胞器结构损伤与渗透压稳态失衡,盐生植物则通过盐分介导的渗透调节机制维持细胞内离子稳态,这一特性赋予其对重金属胁迫的交叉耐受性。相较于非盐生植物(glycophytes),盐生植物利用无机离子(如Na+、Cl-)高效调控胞内渗透平衡的能力显著增强,此为盐生植物应对金属胁迫的独特优势。例如,在盐地碱蓬中,NaCl预处理可抑制镉离子的根部吸收,并通过液泡区室化促进已吸收镉离子的固定[52];浮萍中钠离子转运蛋白基因NHX1的过表达可显著提升其对隔离子胁迫的耐受性[53]。植物对胁迫的响应并非单一模式,可大致分为两类:一是引发直接损伤的急性应激反应;二是诱导胁迫记忆形成的适应性反应。该记忆可使植物再次遭遇胁迫时做出更快、更强的响应,从而提升耐受性[54]。在表达出胁迫之后,植物会拥有一种提高植物应激的胁迫记忆[55]。
与动物类似,预先暴露于应激条件的植物会增强对随后施加的应激的耐受性。在滨海卡克勒(Cakile maritime)中,盐度预处理可通过激活CAT、SOD等抗氧化酶系统,显著提升其对氧化胁迫的耐受性[54]。这种胁迫记忆可维持数周甚至更长时间,使预暴露植物比未暴露植物更高效地抵御活性氧损伤[56]。在胁迫记忆中,水杨酸(SA)、茉莉酸(JA)和脱落酸(ABA)等激素也起着核心调控作用:盐生植物滨海卡克勒暴露在高盐浓度时,JA合成通路被激活,JA浓度升高可诱导抗氧化系统增强,使过氧化氢(H2O2)和丙二醛(MDA,脂质过氧化产物)水平维持在较低水平[57]。而胁迫预处理(如低盐预暴露)可通过负反馈调节降低后续盐胁迫下叶片中的JA水平,同时减轻氧化应激反应[58]。此外,干旱和镉胁迫引发的代谢响应,与盐胁迫预暴露诱导的代谢启动状态具有相似性,表明植物可能通过共享的ABA信号通路实现高效的跨胁启动机制[59]。值得注意的是,植物能够通过感知当前环境信号进而激活调控适应性形态、利于后续适应的基因表达,此现象称为“环境预期”。其与“胁迫记忆”二者在植物胁迫响应调控中具重要意义。深入解析交叉耐受性及相关的胁迫记忆、环境预期等机制,不仅有助于阐明盐生植物的抗逆原理,更可转化为指导抗逆育种的策略。
-
转录组学能系统解析特定生理条件下转录本的分类、定量及调控网络[60],是阐明植物逆境适应分子机制的重要手段。目前盐生植物的转录组学研究已初步展开,但重金属耐受性的分子解析仍存局限,相关研究多停留在生理生化层面,基因功能研究较为匮乏,仅涉及PCs、MT、质膜H+-ATP酶(AHA)、吡咯啉-5-羧酸合成酶(P5CS)及过氧化氢酶(CAT)等少数基因。当前学界争议焦点在于盐生植物对不同非生物胁迫(如盐害与重金属)可能共享相似耐受机制。例如,海马齿经盐处理后对镉的耐受性显著增强;冰叶日中花兼具盐与金属胁迫抗性。两类胁迫均会引发活性氧过量积累、渗透调节物质合成及抗氧化系统激活等共性响应[61]。
现有转录组学研究覆盖盐生植物重金属胁迫和盐胁迫两大方向,结合表1、表2 的比较分析,可鉴定出多胁迫下持续上调的“核心耐受基因”(如 MT、PCs、SOD 等)。这些基因是解析盐-重金属交叉耐受机制的关键,也是分子育种的宝贵资源,借助分子标记辅助选择可快速筛选优良种质,通过基因工程手段编辑或过表达核心基因,可实现目标性状的精准改良。
表 1 重金属胁迫下盐生植物基因表达模式研究
Table 1. Gene expression patterns in saline plants under heavy metal stress.
盐生植物
Halophytes处理
Treatment上调的基因
Up-regulated genes参考文献
References马绊草
Aeluropus littoralis银,汞,铅
Ag, Hg, PbPM H+-ATPase [62] 黑海榄雌
Avicennia germinans铜-镉
Cu-CdAvPCs [29] 海榄雌
Avicennia marina锌,铜,铅,锰
Zn, Cu, Pb, MnAmMT2, AmSOD1, AmSOD2 [34] 木榄
Bruguiera gymnorrhiza锌,铜,铅
Zn, Cu, PbBgMT2 [34] 海雀稗
Paspalum vaginatum镉
CdPCS1, PCS2, CYP450, HSFA4a, UGP [63] 海蓬子
Salvinia biloba锌,铜
Zn, CuSbMT-2 [36] 盐地碱蓬
Suaeda salsa镉
CdPCs, CAT2 [64] 刚毛柽柳
Tamarix hispida镉
CdThDRE1A [65] 罗布麻
Apocynum venetum镉
Cdcytokinin-related DEGs, ROS scavenger genes, CSL, D-GRL [66] 表 2 盐生植物(盐胁迫后)转录组学研究揭示的与重金属适应有交互作用的相关基因
Table 2. Genes associated with toxic heavy metal tolerance revealed by transcriptomic studies in saline plants
盐生植物
Halophytes测序平台
Sequencing platforms上调的与交互耐受性有关基因
Up-regulated genes associated with cross-resistance参考文献
References印度红树
Xylocarpus moluccensisIllumina HiSeq 2000 ERE MYB, bZIP, Cadmium ion responsive genes [72] 柠条锦鸡儿
Caragana korshinskiiIllumina HiSeq 2000 SOD, CATAPXPOX, MYBNAC, ERF [71] 盐生草
Halogeton glomeratusIllumina HiSeq 2000 POD, GPX, ABA responsive genes [68] 假厚藤
Ipomoea imperatiIllumina HiSeq 2500 MYB, HD-ZIP, ABA signaling [69] 花花柴
Karelinia caspicaIllumina HiSeq 2000 ABA responsive genes [67] 冰叶日中花
Mesembryanthemum crystallinumIllumina Genome Analyzer llx P5CS1 [73] 野生盐生稻
Wild Halophyte RiceIllumina Genome Analyzer ll MYB, AP2-EREBP, bZIP, NAC [70] 长叶红砂/黄叶红砂
Reaumuria trigynaIllumina HiSeq 2000 GPX, APX, PODs, SODs [74] 互花米草
Spartina alterniflora LoiselRoche's 454 GS-FLX ARF, MYB, H+-ATPase, vacuolar H+-ATPase, CDPK [75] 盐地鼠尾粟
Sporobolus virginicusIllumina HiSeq 2500 P5CS, ERF, bZIP, MYB, NAC [76] 马来碱蓬
Suaeda fruticosaIllumina HiSeq 2000 APX, cadmium resistance 2-like, Aluminum-activated malate transporter 10, Magnesium transporter NIPA2, Vacuolar lron transporter family [77] 裸花碱蓬
Suaeda maritimeIllumina HiSeq 2000 Cu/Zn-SODMDHAR, Mn-SOD, Fe-SOD [78] 碱蓬
Suaeda glaucaIllumina HiSeq 2500 Oligopeptide transporters, APX, POX [79] 大量研究表明,在盐生植物花花柴(Karelinia caspica)、盐生草(Halogeton glomeratus)及假厚藤(Ipomoea pes-caprae)中,与脱落酸信号通路相关的基因表达显著上调[67-69]。其他盐生植物如野生稻(Oryza rufipogon)、柠条锦鸡儿(Caragana korshinskii)、印度红树(Xylocarpus moluccensis)等物种的MYB、AP2/ERF、bZIP、NAC等转录因子家族,可通过与下游靶基因启动子区的顺式作用元件结合,从而激活胁迫耐受机制[69-72]。WRKY、bHLH等转录因子家族也在盐胁迫信号转导网络中发挥核心调控作用[74]。尽管上述研究为解析盐生植物抗逆机制提供了丰富的抗逆基因资源,但其在重金属胁迫下的转录调控网络构建与验证仍需进行系统探究。
-
土壤微生物通过生物地球化学循环参与盐生植物对重金属的耐受过程:节杆菌属(Arthrobacter)通过分泌草酸、柠檬酸等有机酸酸化土壤以活化重金属;芽孢杆菌属(Bacillus)通过产生胞外多糖(EPS)和生物表面活性剂,高效吸附重金属;假单胞菌属(Pseudomonas)通过Fe3+还原等氧化还原反应,实现重金属转化与钝化;根瘤菌属(Rhizobium)作为固氮菌,其促生特性、结瘤过程及固氮酶活性通常对重金属胁迫敏感,但部分耐金属根瘤菌株已被证实可缓解重金属胁迫,尤其在豆科植物共生体系中[81]。子囊菌门(Ascomycota)和担子菌门(Basidiomycota)真菌可定殖于重金属污染土壤[82]。而丛枝菌根真菌(Arbuscular Mycorrhizal Fungi,AMF)亦广泛分布于重金属污染及贫瘠土壤中[83]。近年来,研究进一步细化了微生物对重金属的生物转化与固定机制。例如,特定细菌和真菌可通过其铬(Cr)还原酶系统将高毒性的Cr(VI)还原为低毒的Cr(III);微生物介导的汞甲基化及砷的价态转化与外排过程,也显著调控重金属的环境迁移性与生态风险[84]。
微生物通过调控金属形态转化、溶解、迁移及毒性缓解等关键过程,与宿主植物形成紧密互作[85]。例如,盐沼植物海米草(Spartina alterniflora)通过根系分泌物吸引硫酸盐还原菌(如Desulfovibrio spp.)定殖,协同增强金属耐受性[86-87],其根系活动显著影响酚氧化酶及磷酸水解酶等胞外酶活性[88],这些酶活性与有机质循环及金属形态转化密切相关。重金属吸收模式呈现物种及形态依赖性:互花米草和狐米草(Spartina patens)对不同形态砷(As)的吸收与转运存在差异,无机砷[As(III)/As(V)]主要滞留于根部(占比>85%),而有机砷(如MMA、DMA)可快速向茎部转运[89]。植物与微生物形成基于代谢交换的协同解毒网络:植物根系分泌物为微生物提供碳源,微生物则通过代谢反馈调控金属有效性。研究发现,微生物不仅能改善根际环境,更能深度调控植物内源的解毒与耐受通路[90-91]。例如,微生物共生可系统上调宿主PCs、MTs及液泡膜转运蛋白(如HMA3、MTP1)的基因表达,促进重金属区室化储存;同时,还能显著激活植物的抗氧化酶系统,有效缓解氧化损伤[92]。金属转化微生物(Metal-transforming microorganisms,MTM)通过调节植物生长周期及土壤生化特性减轻重金属毒性,其作用机制包括氧化还原、酸化、沉淀及络合反应[93]。MTM释放出乙酸(AcOH)、葡萄糖(Glc)、草酸(OA)、苹果酸(MA)等有机酸,降低土壤的 pH,随后隔离可溶性金属离子[94]。除了MTM,其他类型的微生物也通过类似的机制影响土壤金属的生物利用度,例如白刺稻与菌根真菌共生,菌根分泌草酸和柠檬酸(CA),对 Cd2+、Cu2+、Pb2+、Zn2+进行增溶[95]。在木腐菌类中例如多孔菌类和灵芝属可以生产草酸晶体,并将重金属污染物转换成毒性较小的形式,如硫酸锌、硫酸铜、硫酸镉、硝酸铅分别转化成脱水草酸锌、水合草酸铜、三水合草酸镉、草酸铅。重金属胁迫下,植物根系分泌氨基酸、有机酸及植物螯合肽等物质形成金属-配体复合物,为微生物提供碳源;微生物则通过代谢活动反馈增强金属生物有效性,形成互利共生。AMF可显著提升盐生植物对重金属及盐胁迫的耐受性,其机制涉及胁迫信号感知、离子转运蛋白调控及基因表达重编程[96]。近期研究表明,AMF与生物有机肥(Bioorganic Fertilizer,BOF)联用可协同改善重金属污染盐渍土,BOF-AMF联合处理不仅增强植物抗逆性,还通过重塑根际微生物群落,显著提高土壤生物活性与修复效率[97]。
各类根际微生物通过多样化生物地球化学过程,与盐生植物构建紧密互作的协同网络,这一互利共生系统是驱动修复能力提升的核心生物因素。特定促生菌与菌根真菌不仅可直接钝化重金属、促进植物营养吸收,还能系统激活植物的整体抗逆性。因此,在育种与应用实践中,应超越对植物个体的单一改良,转而注重筛选和构建高效的“植物-微生物”修复联合体,通过精准调控根际微环境,实现修复功能的协同增强。
-
盐生植物在长期适应盐碱生境的过程中,演化出高度耦合的多层次交叉耐受机制,为阐明植物复合胁迫应对策略提供了独特视角。现有研究表明,其耐盐系统与重金属适应网络在离子转运(如HMA/CDF蛋白协同)、信号调控(MAPK-ABA/JA通路交叉对话)及表观遗传记忆(DNA甲基化修饰)三大层面存在深度功能耦合。这种多维度协同机制不仅阐明了极端生境植物的生存智慧,也为构建“以盐治毒”的植物修复理论框架奠定了重要基础,其中所揭示的关键基因与通路更是分子设计育种的核心靶标。尽管相关研究已取得显著进展,但盐生植物复合胁迫动态响应网络仍有诸多关键科学问题亟待解答:盐离子如何通过调控SOS1、HKT1等关键转运蛋白的构象与活性,进而影响重金属的跨膜运输与亚细胞分配;ROS信号枢纽与螯合代谢通路的时空协同规律如何实现跨胁迫资源优化配置;根际微生物组与宿主植物间基于代谢物交换的分子对话机制是否构成交叉耐受的“第四重维度”。这些问题的解答需突破传统单组学研究范式,整合空间代谢组学、单细胞表观遗传分析等前沿技术,在分子-细胞-生态多尺度构建动态调控模型。
基于现有机制认知,未来研究应推动从“机制解析”向“定向设计”的跨越。以已鉴定的关键耐受基因(如MT、HMA及各类胁迫响应转录因子)为基础,建立高效的分子标记辅助选择体系,实现对耐逆性与富集性状的快速筛选。在此基础上,结合CRISPR/Cas9等基因编辑技术,对ZIP、Nramp等关键转运系统及螯合基因进行定向改造,创制兼具强耐盐性与高效重金属富集能力的“设计型”修复植物。最终,需突破单一植物改良的范式,将AMF、PGPR等根际益生微生物纳入植物修复体系,通过合成生物学理念构建稳定、高效的“植物-微生物”协同修复模块,以实现重金属原位生物矿化与盐分调控的协同调控。应用层面,盐生植物凭借固有耐逆性、边缘土地的适应性及环境友好型修复特征,具备突出的生态修复潜力。其耐盐特性可利用微咸水或再生水进行灌溉,且根际过程能协同提升重金属生物有效性[98-99]。已有研究证实,海滨锦葵(Kosteletzkya virginica)根系对Pb2+的吸附容量可达68.2 mg/g[100];海马齿可高效富集皮革厂废水中的多种重金属;碱蓬、柽柳等物种在模拟修复中镉去除率超过75%[101]。这些案例为田间工程化应用提供了实证依据。且中国滨海盐碱区与重金属污染带的空间重叠特征,为开展盐生植物修复技术的规模化验证提供了天然试验场。
因此,深度融合分子机制解析与现代育种技术,充分挖掘盐生植物交叉耐受潜力,不仅有助于推动边际土壤的“去毒增绿”与生态恢复,更有望催生盐碱农业与污染治理协同发展的新业态,为全球土地资源的可持续管理贡献创新方案。
A review of the cross-tolerance mechanisms of halophytes to heavy metal pollution and prospects for their remediation application
-
摘要: 盐生植物作为盐碱地生态系统的优势类群,其耐盐机制已在生理、生化、分子和代谢组等不同水平得到系统解析。近年来,土壤重金属-盐碱复合污染问题日益严峻,盐生植物因其对多重胁迫的交叉耐受特性备受关注。研究表明,盐生植物通过重金属酶(HMA)等特异性离子转运系统精准调控重金属的吸收、区室化与外排,并借助抗氧化防御、螯合作用以及与根际微生物互作实现协同解毒。尤为重要的是,盐度与重金属胁迫共享MAPK信号级联、脱落酸/茉莉酸等激素通路及表观遗传调控机制,构成交叉耐受的分子基础。因此,盐生植物不仅能够适应高盐环境,更具备对土壤重金属的富集或稳定能力,成为同步治理盐碱化与重金属污染的绿色方案。本综述系统总结了盐生植物交叉耐受机制的研究进展,结合典型案例,从修复原理及分子育种策略等方面解析其修复应用潜能,旨在为盐碱地-重金属复合污染的绿色可持续治理提供理论依据。Abstract: As the dominant groups in saline ecosystems,halophytes have been systematically studied in their salt tolerance mechanisms at the physiological, biochemical, molecular and metabolomic levels. In recent years, the issue of combined soil heavy metal-saline-alkali pollution has become increasingly severe, drawing significant attention to halophytes due to their cross-tolerance to multiple stresses. Research indicates that halophytes precisely regulate heavy metal uptake, compartmentalization, and efflux through specific ion transport systems such as HMA. They achieve synergistic detoxification via antioxidant defenses and chelation mechanisms (e.g., plant chelating peptides, metallothionein). It is interesting to note that salinity and heavy metal stresses share epigenetic regulatory mechanisms—including MAPK signaling cascades, ABA/JA hormone pathways, thereby establishing the molecular basis for cross-tolerance. Therefore, halophytes are not only adapted to high-salinity environments but also possess the capacity to accumulate or stabilise heavy metals in soil, presenting a green solution for concurrently addressing salinisation and heavy metal contamination. This paper systematically reviews the mechanisms underlying the cross-tolerance traits of halophytes. Drawing upon representative case studies, it examines their remediation potential from multiple perspectives, including restoration principles and molecular breeding improvement strategies, aiming to provide theoretical foundations for the green and sustainable management of combined saline-alkali and heavy metal pollution.
-
Key words:
- halophytes /
- heavy metal /
- cross-tolerance /
- metal transport /
- phytoremediation
-
表 1 重金属胁迫下盐生植物基因表达模式研究
Table 1 Gene expression patterns in saline plants under heavy metal stress.
盐生植物
Halophytes处理
Treatment上调的基因
Up-regulated genes参考文献
References马绊草
Aeluropus littoralis银,汞,铅
Ag, Hg, PbPM H+-ATPase [62] 黑海榄雌
Avicennia germinans铜-镉
Cu-CdAvPCs [29] 海榄雌
Avicennia marina锌,铜,铅,锰
Zn, Cu, Pb, MnAmMT2, AmSOD1, AmSOD2 [34] 木榄
Bruguiera gymnorrhiza锌,铜,铅
Zn, Cu, PbBgMT2 [34] 海雀稗
Paspalum vaginatum镉
CdPCS1, PCS2, CYP450, HSFA4a, UGP [63] 海蓬子
Salvinia biloba锌,铜
Zn, CuSbMT-2 [36] 盐地碱蓬
Suaeda salsa镉
CdPCs, CAT2 [64] 刚毛柽柳
Tamarix hispida镉
CdThDRE1A [65] 罗布麻
Apocynum venetum镉
Cdcytokinin-related DEGs, ROS scavenger genes, CSL, D-GRL [66] 表 2 盐生植物(盐胁迫后)转录组学研究揭示的与重金属适应有交互作用的相关基因
Table 2 Genes associated with toxic heavy metal tolerance revealed by transcriptomic studies in saline plants
盐生植物
Halophytes测序平台
Sequencing platforms上调的与交互耐受性有关基因
Up-regulated genes associated with cross-resistance参考文献
References印度红树
Xylocarpus moluccensisIllumina HiSeq 2000 ERE MYB, bZIP, Cadmium ion responsive genes [72] 柠条锦鸡儿
Caragana korshinskiiIllumina HiSeq 2000 SOD, CATAPXPOX, MYBNAC, ERF [71] 盐生草
Halogeton glomeratusIllumina HiSeq 2000 POD, GPX, ABA responsive genes [68] 假厚藤
Ipomoea imperatiIllumina HiSeq 2500 MYB, HD-ZIP, ABA signaling [69] 花花柴
Karelinia caspicaIllumina HiSeq 2000 ABA responsive genes [67] 冰叶日中花
Mesembryanthemum crystallinumIllumina Genome Analyzer llx P5CS1 [73] 野生盐生稻
Wild Halophyte RiceIllumina Genome Analyzer ll MYB, AP2-EREBP, bZIP, NAC [70] 长叶红砂/黄叶红砂
Reaumuria trigynaIllumina HiSeq 2000 GPX, APX, PODs, SODs [74] 互花米草
Spartina alterniflora LoiselRoche's 454 GS-FLX ARF, MYB, H+-ATPase, vacuolar H+-ATPase, CDPK [75] 盐地鼠尾粟
Sporobolus virginicusIllumina HiSeq 2500 P5CS, ERF, bZIP, MYB, NAC [76] 马来碱蓬
Suaeda fruticosaIllumina HiSeq 2000 APX, cadmium resistance 2-like, Aluminum-activated malate transporter 10, Magnesium transporter NIPA2, Vacuolar lron transporter family [77] 裸花碱蓬
Suaeda maritimeIllumina HiSeq 2000 Cu/Zn-SODMDHAR, Mn-SOD, Fe-SOD [78] 碱蓬
Suaeda glaucaIllumina HiSeq 2500 Oligopeptide transporters, APX, POX [79] -
[1] 刘小京, 郭凯, 封晓辉, 等. 农业高效利用盐碱地资源探讨[J]. 中国生态农业学报(中英文), 2023, 31(3): 345−353. https://doi.org/10.12357/cjea.20220967 doi: 10.12357/cjea.20220967 [2] 于功霞, 李迎, 刘宏元, 等. 我国盐碱地改良与利用技术研究进展[J]. 黑龙江农业科学, 2025(7): 96−103. https://doi.org/10.11942/j.issn1002-2767.2025.07.0096 doi: 10.11942/j.issn1002-2767.2025.07.0096 [3] 王遵亲. 中国盐渍土[M]. 北京: 科学出版社, 1993. (查阅网上资料, 未找到本条文献页码信息, 请补充) [4] 梁淑敏, 许艳萍, 陈裕, 等. 工业大麻对重金属污染土壤的治理研究进展[J]. 生态学报, 2013, 33(5): 1347−1356. https://doi.org/10.5846/stxb201209231342 doi: 10.5846/stxb201209231342 [5] Chibuike G U, Obiora S C. Heavy metal polluted soils: effect on plants and bioremediation methods [J]. Applied and Environmental Soil Science, 2014, 2014: 752708. https://doi.org/10.1155/2014/752708 doi: 10.1155/2014/752708 [6] Hou D Y, Jia X Y, Wang L W, et al. Global soil pollution by toxic metals threatens agriculture and human health [J]. Science, 2025, 388(6744): 316−321. https://doi.org/10.1126/science.adr5214 doi: 10.1126/science.adr5214 [7] Haider F U, Cai L Q, Coulter J A, et al. Cadmium toxicity in plants: impacts and remediation strategies [J]. Ecotoxicology and Environmental Safety, 2021, 211: 111887. https://doi.org/10.1016/j.ecoenv.2020.111887 doi: 10.1016/j.ecoenv.2020.111887 [8] 王智慧, 牛二利, 高远亮, 等. 浙麦冬HMA基因家族鉴定及镉胁迫响应分析[J]. 生物工程学报, 2025, 41(2): 771−790. https://doi.org/10.13345/j.cjb.240475 doi: 10.13345/j.cjb.240475 [9] Cobbett C, Goldsbrough P. Phytochelatins and metallothioneins: roles in heavy metal detoxification and homeostasis [J]. Annual Review of Plant Biology, 2002, 53: 159−182. https://doi.org/10.1146/annurev.arplant.53.100301.135154 doi: 10.1146/annurev.arplant.53.100301.135154 [10] Macfarlane G R, Pulkownik A, Burchett M D. Accumulation and distribution of heavy metals in the grey mangrove, Avicennia marina (Forsk. )Vierh. : biological indication potential [J]. Environmental Pollution, 2003, 123(1): 139−151. https://doi.org/10.1016/S0269-7491(02)00342-1 doi: 10.1016/S0269-7491(02)00342-1 [11] Wang Y T, Qiu Q, Xin G R, et al. Heavy metal contamination in a vulnerable mangrove swamp in South China [J]. Environmental Monitoring and Assessment, 2013, 185(7): 5775−5787. https://doi.org/10.1007/s10661-012-2983-4 doi: 10.1007/s10661-012-2983-4 [12] Nikalje G C, Nikam T D, Suprasanna P. Looking at halophytic adaptation to high salinity through genomics landscape [J]. Current Genomics, 2017, 18(6): 542−552. https://doi.org/10.2174/1389202918666170228143007 doi: 10.2174/1389202918666170228143007 [13] Christofilopoulos S, Syranidou E, Gkavrou G, et al. The role of halophyte Juncus acutus L. in the remediation of mixed contamination in a hydroponic greenhouse experiment [J]. Journal of Chemical Technology & Biotechnology, 2016, 91(6): 1665−1674. https://doi.org/10.1002/jctb.4939 doi: 10.1002/jctb.4939 [14] Pan X, Chen G P, Shi C, et al. Effects of Zn stress on growth, Zn accumulation, translocation, and subcellular distribution of Spartina alterniflora Loisel [J]. CLEAN-Soil, Air, Water, 2016, 44(5): 579−585. https://doi.org/10.1002/clen.201400288 doi: 10.1002/clen.201400288 [15] Gill S S, Khan N A, Tuteja N. Differential cadmium stress tolerance in five indian mustard (Brassica juncea L. ) cultivars: an evaluation of the role of antioxidant machinery [J]. Plant Signaling & Behavior, 2011, 6(2): 293−300. https://doi.org/10.4161/psb.6.2.15049 doi: 10.4161/psb.6.2.15049 [16] Møller I M, Jensen P E, Hansson A. Oxidative modifications to cellular components in plants [J]. Annual Review of Plant Biology, 2007, 58: 459−481. https://doi.org/10.1146/annurev.arplant.58.032806.103946 doi: 10.1146/annurev.arplant.58.032806.103946 [17] Sharma S S, Dietz K J. The relationship between metal toxicity and cellular redox imbalance [J]. Trends in Plant Science, 2009, 14(1): 43−50. https://doi.org/10.1016/j.tplants.2008.10.007 doi: 10.1016/j.tplants.2008.10.007 [18] Keunen E, Remans T, Bohler S, et al. Metal-induced oxidative stress and plant mitochondria [J]. International Journal of Molecular Sciences, 2011, 12(10): 6894−6918. https://doi.org/10.3390/ijms12106894 doi: 10.3390/ijms12106894 [19] Tsukagoshi H, Busch W, Benfey P N. Transcriptional regulation of ROS controls transition from proliferation to differentiation in the root [J]. Cell, 2010, 143(4): 606−616. https://doi.org/10.1016/j.cell.2010.10.020 doi: 10.1016/j.cell.2010.10.020 [20] Rucińiska-Sobkowiak R. Oxidative stress in plants exposed to heavy metals [J]. Postepy Biochemii, 2010, 56(2): 191−200. https://doi.org/10.1201/9781003022879-7 doi: 10.1201/9781003022879-7 [21] Miller G, Suzuki N, Ciftci-Yilmaz S, et al. Reactive oxygen species homeostasis and signalling during drought and salinity stresses [J]. Plant, Cell & Environment, 2010, 33(4): 453−467. https://doi.org/10.1111/j.1365-3040.2009.02041.x doi: 10.1111/j.1365-3040.2009.02041.x [22] 王会方, 於朝广, 王涛, 等. 硅缓解植物重金属毒害机理的研究进展[J]. 云南农业大学学报(自然科学), 2016, 31(3): 528−535. https://doi.org/10.16211/j.issn.1004-390X(n).2016.03.021 doi: 10.16211/j.issn.1004-390X(n).2016.03.021 [23] 郑昕煜, 王辰雨, 周益峰, 等. 金属伴侣蛋白在植物重金属解毒中的作用[J]. 植物生理学报, 2025, 61(7): 916−925. https://doi.org/10.13592/j.cnki.ppj.300310 doi: 10.13592/j.cnki.ppj.300310 [24] 陈沁. 氯化血红素通过HO1/CO、Fe2+途径参与缓解水稻幼苗重金属胁迫[D]. 南京: 南京农业大学, 2016. [25] 卢诗雨, 令建伟, 吴琳, 等. 湿地植物对重金属污染的去除机制研究进展[J]. 湿地科学, 2025, 23(4): 846−857. https://doi.org/10.13248/j.cnki.wetlandsci.20240183 doi: 10.13248/j.cnki.wetlandsci.20240183 [26] Anjum N A, Hasanuzzaman M, Hossain M A, et al. Jacks of metal/metalloid chelation trade in plants-an overview [J]. Frontiers in Plant Science, 2015, 6: 192. https://doi.org/10.3389/fpls.2015.00192 doi: 10.3389/fpls.2015.00192 [27] 杨柳. 刚毛藻植物螯合肽对Pb胁迫的响应及吸收[D]. 合肥: 安徽农业大学, 2022. https://doi.org/10.26919/d.cnki.gannu.2022.000791 [28] Tennstedt P, Peisker D, Böttcher C, et al. Phytochelatin synthesis is essential for the detoxification of excess zinc and contributes significantly to the accumulation of zinc [J]. Plant Physiology, 2009, 149(2): 938−948. https://doi.org/10.1104/pp.108.127472 doi: 10.1104/pp.108.127472 [29] Gonzalez-Mendoza D, Moreno A Q, Zapata-Perez O. Coordinated responses of phytochelatin synthase and metallothionein genes in black mangrove, Avicennia germinans, exposed to cadmium and copper [J]. Aquatic Toxicology, 2007, 83(4): 306−314. https://doi.org/10.1016/j.aquatox.2007.05.005 doi: 10.1016/j.aquatox.2007.05.005 [30] Bankaji I, Caçador I, Sleimi N. Physiological and biochemical responses of Suaeda fruticosa to cadmium and copper stresses: growth, nutrient uptake, antioxidant enzymes, phytochelatin, and glutathione levels [J]. Environmental Science and Pollution Research, 2015, 22(17): 13058−13069. https://doi.org/10.1007/s11356-015-4414-x doi: 10.1007/s11356-015-4414-x [31] Cobbett C, Goldsbrough P. Phytochelatins and metallothioneins: roles in heavy metal detoxification and homeostasis [J]. Annual Review of Plant Biology, 2002, 53: 159−182. (查阅网上资料,本条文献与第9条文献重复,请确认) https://doi.org/10.1146/annurev.arplant.53.100301.135154 [32] Guo W J, Bundithya W, Goldsbrough P B. Characterization of the Arabidopsis metallothionein gene family: tissue-specific expression and induction during senescence and in response to copper [J]. New Phytologist, 2003, 159(2): 369−381. https://doi.org/10.1046/j.1469-8137.2003.00813.x doi: 10.1046/j.1469-8137.2003.00813.x [33] Pakdee O, Tshering S, Pokethitiyook P, et al. Examination of the metallothionein gene family in greater duckweed Spirodela polyrhiza [J]. Plants, 2022, 12(1): 125. https://doi.org/10.3390/plants12010125 doi: 10.3390/plants12010125 [34] Huang G Y, Wang Y S. Expression analysis of type 2 metallothionein gene in mangrove species (Bruguiera gymnorrhiza) under heavy metal stress [J]. Chemosphere, 2009, 77(7): 1026−1029. https://doi.org/10.1016/j.chemosphere.2009.07.073 doi: 10.1016/j.chemosphere.2009.07.073 [35] Usha B, Venkataraman G, Parida A. Heavy metal and abiotic stress inducible metallothionein isoforms from Prosopis juliflora (SW) D. C. show differences in binding to heavy metals in vitro [J]. Molecular Genetics and Genomics, 2009, 281(1): 99−108. https://doi.org/10.1007/s00438-008-0398-2 doi: 10.1007/s00438-008-0398-2 [36] 赵之伟, 曹冠华, 李涛. 金属硫蛋白的研究进展[J]. 云南大学学报(自然科学版), 2013, 35(3): 390−398. https://doi.org/10.7540/j.ynu.20130187 doi: 10.7540/j.ynu.20130187 [37] Yang J L, Wang Y C, Liu G F, et al. Tamarix hispida metallothionein-like ThMT3, a reactive oxygen species scavenger, increases tolerance against Cd2+, Zn2+, Cu2+, and NaCl in transgenic yeast [J]. Molecular Biology Reports, 2011, 38(3): 1567−1574. https://doi.org/10.1007/s11033-010-0265-1 doi: 10.1007/s11033-010-0265-1 [38] Sun X H, Yu G, Li J T, et al. A heavy metal-associated protein (AcHMA1) from thehalophyte, Atriplex canescens (Pursh) Nutt. , confers tolerance to iron and other abioticstresses when expressed in Saccharomyces cerevisiae [J]. International Journal of Molecular Sciences, 2014, 15(8): 14891-14906. https://doi.org/10.3390/ijms150814891 [39] Su H, Balderas E, Vera-Estrella R, et al. Expression of the cation transporter McHKT1 in a halophyte [J]. Plant Molecular Biology, 2003, 52(5): 967−980. https://doi.org/10.1023/a:1025445612244 doi: 10.1023/a:1025445612244 [40] Papoyan A, Kochian L V. Identification of Thlaspi caerulescens genes that may be involved in heavy metal hyperaccumulation and tolerance. Characterization of a novel heavy metal transporting ATPase [J]. Plant Physiology, 2004, 136(3): 3814−3823. https://doi.org/10.1104/pp.104.044503 doi: 10.1104/pp.104.044503 [41] Belouchi A, Kwan T, Gros P. Cloning and characterization of the OsNramp family from Oryza sativa, a new family of membrane proteins possibly implicated in the transport of metal ions [J]. Plant Molecular Biology, 1997, 33(6): 1085−1092. https://doi.org/10.1023/A:1005723304911 doi: 10.1023/A:1005723304911 [42] 刘霄霏, 甘锦鑫, 周涛, 等. 千穗谷MTP基因家族生信分析及镉胁迫下的表达特征[J]. 植物科学学报, 2023, 41(4): 467−478. https://doi.org/10.11913/PSJ.2095-0837.22279 doi: 10.11913/PSJ.2095-0837.22279 [43] Van Der Zaal B J, Neuteboom L W, Pinas J E, et al. Overexpression of a novel Arabidopsis gene related to putative zinc-transporter genes from animals can lead to enhanced zinc resistance and accumulation [J]. Plant Physiology, 1999, 119(3): 1047−1056. https://doi.org/10.1104/pp.119.3.1047 doi: 10.1104/pp.119.3.1047 [44] Llorens E, González-Hernández A I, Scalschi L, et al. Priming mediated stress and cross-stress tolerance in plants: concepts and opportunities[M]//Hossain M A, Liu F L, Burritt D J, et al. Priming-mediated stress and cross-stress tolerance in crop plants. London: Academic Press, 2020: 1−20. https://doi.org/10.1016/B978-0-12-817892-8.00001-5 [45] Foyer C H, Rasool B, Davey J W, et al. Cross-tolerance to biotic and abiotic stresses in plants: a focus on resistance to aphid infestation [J]. Journal of Experimental Botany, 2016, 67(7): 2025−2037. https://doi.org/10.1093/jxb/erw079 doi: 10.1093/jxb/erw079 [46] Dhar R, Sägesser R, Weikert C, et al. Yeast adapts to a changing stressful environment by evolving cross-protection and anticipatory gene regulation [J]. Molecular Biology and Evolution, 2013, 30(3): 573−588. https://doi.org/10.1093/molbev/mss253 doi: 10.1093/molbev/mss253 [47] Knight H, Knight M R. Abiotic stress signalling pathways: specificity and cross-talk [J]. Trends in Plant Science, 2001, 6(6): 262−267. https://doi.org/10.1016/s1360-1385(01)01946-x doi: 10.1016/s1360-1385(01)01946-x [48] Capiati D A, País S M, Téllez-Iñón M T. Wounding increases salt tolerance in tomato plants: evidence on the participation of calmodulin-like activities in cross-tolerance signalling [J]. Journal of Experimental Botany, 2006, 57(10): 2391−2400. https://doi.org/10.1093/jxb/erj212 doi: 10.1093/jxb/erj212 [49] Chinnusamy V, Schumaker K, Zhu J K. Molecular genetic perspectives on cross-talk and specificity in abiotic stress signalling in plants [J]. Journal of Experimental Botany, 2004, 55(395): 225−236. https://doi.org/10.1093/jxb/erh005 doi: 10.1093/jxb/erh005 [50] Munné-Bosch S, Queval G, Foyer C H. The impact of global change factors on redox signaling underpinning stress tolerance [J]. Plant Physiology, 2013, 161(1): 5−19. https://doi.org/10.1104/pp.112.205690 doi: 10.1104/pp.112.205690 [51] Amtmann A. Learning from evolution: thellungiella generates new knowledge on essential and critical components of abiotic stress tolerance in plants [J]. Molecular Plant, 2009, 2(1): 3−12. https://doi.org/10.1093/mp/ssn094 doi: 10.1093/mp/ssn094 [52] 刘岚钰. NaCl缓解盐地碱蓬Cd毒性机制研究[D]. 西安: 陕西科技大学, 2022. https://doi.org/10.27290/d.cnki.gxbqc.2022.000265 [53] 杨琳. Na+/H+逆向转运蛋白提高浮萍镉胁迫抗性的机制研究[D]. 天津: 天津师范大学, 2017. [54] Walter J, Jentsch A, Beierkuhnlein C, et al. Ecological stress memory and cross stress tolerance in plants in the face of climate extremes [J]. Environmental and Experimental Botany, 2013, 94: 3−8. https://doi.org/10.1016/j.envexpbot.2012.02.009 doi: 10.1016/j.envexpbot.2012.02.009 [55] Bruce T J A, Matthes M C, Napier J A, et al. Stressful “memories” of plants: evidence and possible mechanisms [J]. Plant Science, 2007, 173(6): 603−608. https://doi.org/10.1016/j.plantsci.2007.09.002 doi: 10.1016/j.plantsci.2007.09.002 [56] Ellouzi H, Hamed K B, Cela J, et al. Early effects of salt stress on the physiological and oxidative status of Cakile maritima (halophyte) and Arabidopsis thaliana (glycophyte) [J]. Physiologia Plantarum, 2011, 142(2): 128−143. https://doi.org/10.1111/j.1399-3054.2011.01450.x doi: 10.1111/j.1399-3054.2011.01450.x [57] Jaskiewicz M, Conrath U, Peterhänsel C. Chromatin modification acts as a memory for systemic acquired resistance in the plant stress response [J]. The EMBO Reports, 2011, 12(1): 50−55. https://doi.org/10.1038/embor.2010.186 doi: 10.1038/embor.2010.186 [58] Ryals J A, Neuenschwander U H, Willits M G, et al. Systemic acquired resistance [J]. The Plant Cell, 1996, 8(10): 1809−1819. https://doi.org/10.1105/tpc.8.10.1809 doi: 10.1105/tpc.8.10.1809 [59] Conrath U. Chapter 9 priming of induced plant defense responses [J]. Advances in Botanical Research, 2009, 51: 361−395. https://doi.org/10.1016/S0065-2296(09)51009-9 doi: 10.1016/S0065-2296(09)51009-9 [60] Wang Z, Gerstein M, Snyder M. RNA-Seq: a revolutionary tool for transcriptomics [J]. Nature Reviews Genetics, 2009, 10(1): 57−63. https://doi.org/10.1038/nrg2484 doi: 10.1038/nrg2484 [61] Sruthi P, Shackira A M, Puthur J T. Heavy metal detoxification mechanisms in halophytes: an overview [J]. Wetlands Ecology and Management, 2017, 25(2): 129−148. https://doi.org/10.1007/s11273-016-9513-z doi: 10.1007/s11273-016-9513-z [62] Jam M, Alemzadeh A, Tale A M, et al. Heavy metal regulation of plasma membrane H+-ATPase gene expression in halophyte Aeluropus littoralis [J]. Molecular Biology Research Communications, 2014, 3(2): 129−139. [63] Chen Y, Chen C M, Tan Z Q, et al. Functional identification and characterization of genes cloned from halophyte seashore paspalum conferring salinity and cadmium tolerance [J]. Frontiers in Plant Science, 2016, 7: 102. https://doi.org/10.3389/fpls.2016.00102 doi: 10.3389/fpls.2016.00102 [64] Cong M, Lv J S, Liu X L, et al. Gene expression responses in Suaeda salsa after cadmium exposure [J]. SpringerPlus, 2013, 2(1): 232. https://doi.org/10.1186/2193-1801-2-232 doi: 10.1186/2193-1801-2-232 [65] Xie Q J, Wang D N, Ding Y T, et al. The ethylene response factor gene, ThDRE1A, is involved in abscisic acid- and ethylene-mediated cadmium accumulation in Tamarix hispida [J]. Science of the Total Environment, 2024, 937: 173422. https://doi.org/10.1016/j.scitotenv.2024.173422 doi: 10.1016/j.scitotenv.2024.173422 [66] Jing C L, Wang M, Lu X L, et al. Transcriptome analysis reveals how cadmium promotes root development and accumulates in Apocynum venetum, a promising plant for greening cadmium-contaminated soil [J]. Ecotoxicology and Environmental Safety, 2024, 270: 115872. https://doi.org/10.1016/j.ecoenv.2023.115872 doi: 10.1016/j.ecoenv.2023.115872 [67] Zhang X, Liao M S, Chang D, et al. Comparative transcriptome analysis of the Asteraceae halophyte Karelinia caspica under salt stress [J]. BMC Research Notes, 2014, 7: 927. https://doi.org/10.1186/1756-0500-7-927 doi: 10.1186/1756-0500-7-927 [68] Wang J C, Meng Y X, LI B C, et al. Physiological and proteomic analyses of salt stress response in the halophyte Halogeton glomeratus [J]. Plant, Cell & Environment, 2015, 38(4): 655−669. https://doi.org/10.1111/pce.12428 doi: 10.1111/pce.12428 [69] Luo Y, Reid R, Freese D, et al. Salt tolerance response revealed by RNA-Seq in a diploid halophytic wild relative of sweet potato [J]. Scientific Reports, 2017, 7(1): 9624. https://doi.org/10.1038/s41598-017-09241-x doi: 10.1038/s41598-017-09241-x [70] Garg R, Verma M, Agrawal S, et al. Deep transcriptome sequencing of wild halophyte rice, Porteresia coarctata, provides novel insights into the salinity and submergence tolerance factors [J]. DNA Research, 2014, 21(1): 69−84. https://doi.org/10.1093/dnares/dst042 doi: 10.1093/dnares/dst042 [71] Li S F, Fan C M, Li Y, et al. Effects of drought and salt-stresses on gene expression in Caragana korshinskii seedlings revealed by RNA-seq [J]. BMC Genomics, 2016, 17: 200. https://doi.org/10.1186/s12864-016-2562-0 doi: 10.1186/s12864-016-2562-0 [72] Krishnamurthy P, Mohanty B, Wijaya E, et al. Transcriptomics analysis of salt stress tolerance in the roots of the mangrove Avicennia officinalis [J]. Scientific Reports, 2017, 7(1): 10031. https://doi.org/10.1038/s41598-017-10730-2 doi: 10.1038/s41598-017-10730-2 [73] Tsukagoshi H, Suzuki T, Nishikawa K, et al. RNA-Seq analysis of the response of the halophyte, Mesembryanthemum crystallinum (ice plant) to high salinity [J]. PLoS One, 2015, 10(2): e0118339. https://doi.org/10.1371/journal.pone.0118339 doi: 10.1371/journal.pone.0118339 [74] Dang Z H, Zheng L L, Wang J, et al. Transcriptomic profiling of the salt-stress response in the wild recretohalophyte Reaumuria trigyna [J]. BMC Genomics, 2013, 14: 29. https://doi.org/10.1186/1471-2164-14-29 doi: 10.1186/1471-2164-14-29 [75] Bedre R, Mangu V R, Srivastava S, et al. Transcriptome analysis of smooth cordgrass (Spartina alterniflora Loisel), a monocot halophyte, reveals candidate genes involved in its adaptation to salinity [J]. BMC Genomics, 2016, 17(1): 657. https://doi.org/10.1186/s12864-016-3017-3 doi: 10.1186/s12864-016-3017-3 [76] Yamamoto N, Takano T, Tanaka K, et al. Comprehensive analysis of transcriptome response to salinity stress in the halophytic turf grass Sporobolus virginicus [J]. Frontiers in Plant Science, 2015, 6: 241. https://doi.org/10.3389/fpls.2015.00241 doi: 10.3389/fpls.2015.00241 [77] Diray-Arce J, Clement M, Gul B, et al. Transcriptome assembly, profiling and differential gene expression analysis of the halophyte Suaeda fruticosa provides insights into salt tolerance [J]. BMC Genomics, 2015, 16(1): 353. https://doi.org/10.1186/s12864-015-1553-x doi: 10.1186/s12864-015-1553-x [78] Gharat S A, Parmar S, Tambat S, et al. Transcriptome analysis of the response to NaCl in Suaeda maritima provides an insight into salt tolerance mechanisms in halophytes [J]. PLoS One, 2016, 11(9): e0163485. https://doi.org/10.1371/journal.pone.0163485 doi: 10.1371/journal.pone.0163485 [79] Jin H X, Dong D K, Yang Q H, et al. Salt-responsive transcriptome profiling of Suaeda glauca via RNA sequencing [J]. PLoS One, 2016, 11(3): e0150504. https://doi.org/10.1371/journal.pone.0150504 doi: 10.1371/journal.pone.0150504 [80] Anjum N A, Gill S S, Tuteja N. Enhancing cleanup of environmental pollutants [M]. Cham: Springer, 2017: 275−288. https://doi.org/10.1007/978-3-319-55423-5 [81] Pan L Y, Ma J J, Li J M, et al. Advances of salt stress-responsive transcription factors in plants [J]. Chinese Journal of Biotechnology, 2022, 38(1): 50−65.(查阅网上资料,本条文献是中文文献,请确认) https://doi.org/10.13345/j.cjb.210135 [82] Narendrula-Kotha R, Nkongolo K K. Microbial response to soil liming of damaged ecosystems revealed by pyrosequencing and phospholipid fatty acid analyses [J]. PLoS One, 2017, 12(1): e0168497. https://doi.org/10.1371/journal.pone.0168497 doi: 10.1371/journal.pone.0168497 [83] Khan A G, Kuek C, Chaudhry T M, et al. Role of plants, mycorrhizae and phytochelators in heavy metal contaminated land remediation [J]. Chemosphere, 2000, 41(1/2): 197−207. https://doi.org/10.1016/S0045-6535(99)00412-9 doi: 10.1016/S0045-6535(99)00412-9 [84] Yu Y, Li Z M, Liu Y H, et al. Roles of plant-associated microorganisms in regulating the fate of Hg in croplands: a perspective on potential pathways in maintaining sustainable agriculture [J]. Science of the Total Environment, 2022, 834: 155204. https://doi.org/10.1016/j.scitotenv.2022.155204 doi: 10.1016/j.scitotenv.2022.155204 [85] Kong Z Y, Glick B R. The role of plant growth-promoting bacteria in metal phytoremediation [J]. Advances in Microbial Physiology, 2017, 71: 97−132. https://doi.org/10.1016/bs.ampbs.2017.04.001 doi: 10.1016/bs.ampbs.2017.04.001 [86] Otero X L, Macías F. Spatial and seasonal variation in heavy metals in interstitial water of salt marsh soils [J]. Environmental Pollution, 2002, 120(2): 183−190. https://doi.org/10.1016/S0269-7491(02)00159-8 doi: 10.1016/S0269-7491(02)00159-8 [87] Otero X L, Macias F. Variation with depth and season in metal sulfides in salt marsh soils [J]. Biogeochemistry, 2002, 61(3): 247−268. https://doi.org/10.1023/A:1020230213864 doi: 10.1023/A:1020230213864 [88] Reboreda R, Caçador I. Enzymatic activity in the rhizosphere of Spartina maritima: potential contribution for phytoremediation of metals [J]. Marine Environmental Research, 2008, 65(1): 77−84. https://doi.org/10.1016/j.marenvres.2007.09.001 doi: 10.1016/j.marenvres.2007.09.001 [89] Carbonell A A, Aarabi M A, Delaune R D, et al. Arsenic in wetland vegetation: availability, phytotoxicity, uptake and effects on plant growth and nutrition [J]. Science of the Total Environment, 1998, 217(3): 189−199. https://doi.org/10.1016/S0048-9697(98)00195-8 doi: 10.1016/S0048-9697(98)00195-8 [90] Lin S Q, He Q, Zhang M X, et al. Effects of cadmium stress on root exudates and soil rhizosphere microorganisms of rice (Oryza sativa L. ) and its ecological regulatory mechanisms [J]. Plants, 2025, 14(11): 1695. https://doi.org/10.3390/plants14111695 doi: 10.3390/plants14111695 [91] Liu Y, He G D, He T B, et al. Signaling and detoxification strategies in plant-microbes symbiosis under heavy metal stress: a mechanistic understanding [J]. Microorganisms, 2022, 11(1): 69. https://doi.org/10.3390/microorganisms11010069 doi: 10.3390/microorganisms11010069 [92] Sánchez-Thomas R, Hernández-Garnica M, Granados-Rivas J C, et al. Intertwining of cellular osmotic stress handling mechanisms and heavy metal accumulation [J]. Molecular Biotechnology, 2025, 67(12): 4386−4402. https://doi.org/10.1007/s12033-024-01351-y doi: 10.1007/s12033-024-01351-y [93] Singh J S, Seneviratne G. Agro-environmental sustainability: volume 2: managing environmental pollution [M]. Cham: Springer, 2017: 147−163. https://doi.org/10.1007/978-3-319-49727-3 [94] Dighton J, White J F. The fungal community [M]. 4th ed. Boca Raton: CRC Press, 2017: 287−305. https://doi.org/10.1201/9781315119496 [95] Gadd G M, Bahri-Esfahani J, Li Q W, et al. Oxalate production by fungi: significance in geomycology, biodeterioration and bioremediation [J]. Fungal Biology Reviews, 2014, 28(2/3): 36−55. https://doi.org/10.1016/j.fbr.2014.05.001 doi: 10.1016/j.fbr.2014.05.001 [96] Saxena B, Sharma K, Kapoor R, et al. Insights into the molecular aspects of salt stress tolerance in mycorrhizal plants [J]. World Journal of Microbiology and Biotechnology, 2022, 38(12): 253. https://doi.org/10.1007/s11274-022-03440-z doi: 10.1007/s11274-022-03440-z [97] Lu C Y, Zhang Z C, Guo P R, et al. Synergistic mechanisms of bioorganic fertilizer and AMF driving rhizosphere bacterial community to improve phytoremediation efficiency of multiple HMs-contaminated saline soil [J]. Science of the Total Environment, 2023, 883: 163708. https://doi.org/10.1016/j.scitotenv.2023.163708 doi: 10.1016/j.scitotenv.2023.163708 [98] Nikalje G C, Srivastava A K, Pandey G K, et al. Halophytes in biosaline agriculture: mechanism, utilization, and value addition [J]. Land Degradation & Development, 2018, 29(4): 1081−1095. https://doi.org/10.1002/ldr.2819 doi: 10.1002/ldr.2819 [99] Lutts S, Qin P, Han R M. Salinity influences biosorption of heavy metals by the roots of the halophyte plant species Kosteletzkya pentacarpos [J]. Ecological Engineering, 2016, 95: 682−689. https://doi.org/10.1016/j.ecoleng.2016.06.009 doi: 10.1016/j.ecoleng.2016.06.009 [100] Caparrós P G, Ozturk M, Gul A, et al. Halophytes have potential as heavy metal phytoremediators: a comprehensive review [J]. Environmental and Experimental Botany, 2022, 193: 104666. https://doi.org/10.1016/j.envexpbot.2021.104666 doi: 10.1016/j.envexpbot.2021.104666 [101] Ayyappan D, Sathiyaraj G, Ravindran K C. Phytoextraction of heavy metals by Sesuvium portulacastrum l. a salt marsh halophyte from tannery effluent [J]. International Journal of Phytoremediation, 2016, 18(5): 453−459. https://doi.org/10.1080/15226514.2015.1109606 doi: 10.1080/15226514.2015.1109606 -
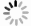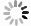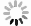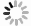
 点击查看大图
点击查看大图
计量
- 文章访问数: 407
- HTML全文浏览量: 202
- 被引次数: 0


 下载:
下载: